 Research Article
Research Article
Genetic Polymorphisms in New Zealand Sheep Breeds
Ugonna J Ekegbu*, Ishaku L Haruna, Ghassan Mahmoud, Huitong Zhou and Jon GH Hickford
Faculty of Agriculture and Life Sciences, New Zealand
Ugonna J Ekegbu, Faculty of Agriculture and Life Sciences, Lincoln 7647, New Zealand.
Received Date:November 06, 2018; Published Date: November 21, 2018
Abstract
Animal production is a booming industry with the improvement of economically desirable traits as its primary concern. Markerassisted selection utilizes genetic variations within candidate genes that influence production traits as a means of guiding animal breeding and improving the traits of interest. Growth hormone (GH) plays a crucial role in pre-natal muscular and bone growth and development. GH brings about various physiological functions either directly by binding its receptor or indirectly by stimulating the release of insulin growth factor 1 (IGF1). Insulin growth factor 1 receptor (IGF1R) mediates its function on metabolism, homeostasis and development upon the binding of IGF1. The POU-domain class 1 transcription factor 1 (POU1F1) regulates the pre-natal development of cells of the anterior pituitary, including somatotrophs that produce GH. Reports have demonstrated associations between polymorphisms in these genes and animal production traits. This novel study examined the polymorphisms in the coding regions of candidate genes, GH2Z, IGF1R and POU1F1, in New Zealand (NZ) sheep. The sheep breeds investigated were NZ Romney and Merino, two commercially sought-after breeds. The results revealed two variants, AA and AB, for the exon 3 of POU1F1. The AA and AB genotypes had frequencies of 78% and 22% for Romney sheep, and 64% and 36% for Merino sheep respectively. All frequencies were in accordance with the Hardy-Weinberg Equilibrium (P > 0.05). The exon 2 of GH2Z revealed multiple variations while no variation was detected for the exon 15 of IGF1R.
Keywords: Marker-assisted selection; Polymorphism; Sheep; Gene
Introduction
The improvement of phenotypic traits such as growth rate, meat quality and milk production is at the core of the animal production industry. These production traits are dependent on a delicate interplay of genes. Research into finding variations in candidate genes that underpin production traits is becoming increasingly popular.
Growth is a complex physiological process involving a vast array of hormones and an interplay of signaling pathways and essential to all living organisms. Growth hormone (GH) or somatotropin is a 191-amino acid peptide hormone secreted by the anterior pituitary gland, which plays a prominent role in the control of growth and metabolism [1]. Ovine growth hormone is about 2.9 kbp long with five exons interspersed by four introns and is located on chromosome 11. Sheep GH differs from that of cattle and human GH but is similar to goat GH, in that it has two duplicate yet distinct alleles GH1 and GH2, the latter which is also a duplicate resulting in GH2-N and GH2-Z [2,3]. GH stimulates the somatic growth and development of essentially all body tissues, as well as regulates metabolism and body composition[2]. GH exerts its physiological effects directly or indirectly.
Insulin growth factor 1 (IGF-1) is a hormone structurally similar to insulin. IGF-1 secretion by the liver is regulated by GH hence IGF-1 serving as a chief mediator of GH effects by binding to its receptor, IGF1R [4]. When activated by IGF-1 or other similar ligand binding, IGF1R plays a crucial role in cell growth and metabolism by initiating a cascade of molecular interactions commonly via phosphotidylinositol-3-kinase/protein kinase B (PI3K-AKT) pathway [5,6]. In sheep, IGF1R gene is located on chromosome 11, comprising of 20 exons and 19 introns.
In sheep, POU1F1 is 17 kbp gene located on sheep chromosome 1, consisting of six exons and five introns and encoding a 33 kDa protein [7]. POU1F1 is a positive regulator of growth hormone, prolactin and thyroid-stimulating hormone β in mammals, and is crucial to the differentiation and survival of three anterior pituitary cell types: thyrotropes, somatotrophs and lactotrophs[8].
The aforementioned genes are all found in the vicinity of the quantitative trait loci (QTL) for economically beneficial traits [9]. The Romney and Merino breeds are two of the most popular breeds in New Zealand, the former bred for both meat and wool, while the latter is solely bred for wool. The meat and wool industry are major drivers of many economies in the world, including New Zealand [10]. The study aims to search for polymorphisms within the GH, IGF1R and POU1F1 genes, which could potentially serve as DNA markers.
Experimental Details
Animals and data collection
All research involving animals was performed in accordance with the Animal Welfare Act 1999 (NZ Government). Three hundred sheep breeds were examined in this study - Romney (n = 150) and Merino (n = 150). The lambs were progeny of non-consanguineous sires. Upon birth, each animal was given an identification number and details surrounding birth like birth weight, birth date and gender were recorded. Blood samples were collected from each lamb onto an FTA card (Whatman Bioscience, Middlesex, UK) by piercing the ears of the sheep. DNA was purified from the blood sample using a method where the disc containing blood sample is hydrolyzed with 200μl of sodium hydroxide at 60 0C, and subsequently rinsed with 200μl of TBE (Tris Borate EDTA) buffer [11].
Materials and primer design
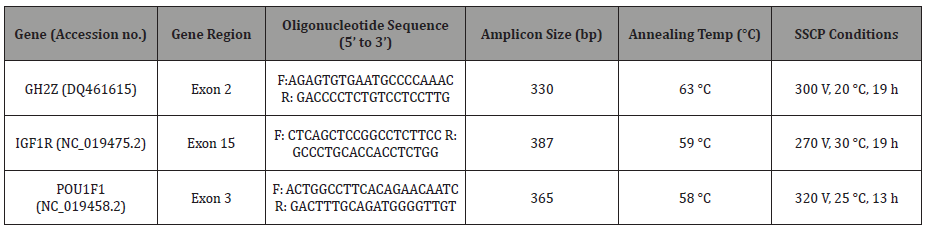
Oligonucleotide primers were designed for the genes, POU1F1, GH2Z and IGF1R based on NCBI database on sheep (Table 1). The primers were manufactured by Integrated DNA Technologies, IA, USA.
Table 1: Primer oligonucleotide sequences, NCBI accession numbers and PCR conditions.

Polymerase chain reaction (PCR) and single-nucleotide conformational polymorphism (SSCP)
PCR was amplified in 15μl reaction tubes containing 1.2 mm punch of the FTA card, 0.25μM of each primer, 0.5 U of Taq DNA polymerase (Qiagen, Hilden, Germany), 150μM of each deoxyribonucleoside triphosphate (dNTP) (Bio line, London, UK), 2.5 mM Mg2+, and deionized water (dH2O) to make up to volume. Bio-Rad S1000 thermal cyclers (Bio-Rad, Hercules, CA, USA) were used to perform DNA amplifications. The amplification conditions included an initial denaturation at 94 °C for 2min, followed by 34 cycles of 94 °C for 30s, annealing for 30s (temperature differs for genes, Table 1), and 72 ºC for 30s, and a final extension step at 72 °C for 5min.
The success of PCR was first confirmed using agarose gel electrophoresis. 1μl of ethidium bromide was added to 1% agarose, with wells into which 2μl of sample mixed with 8μl of bromophenol blue dye was added.
The amplified PCR products were then subjected to SSCP (single-nucleotide conformational polymorphism) analysis. A 0.7μl aliquot of the amplicons was added to 7μl of loading dye containing 10mM EDTA, 0.025% bromophenol blue, 0.025% xylene-cyanole, 98% formamide. Initial denaturation at 95°C for 5min was followed by rapid cooling of the samples on wet ice. The samples were then loaded onto 16 cm X 18 cm, acrylamide: bisacrylamide (37.5:1) (Bio-Rad) gels. Electrophoresis was carried out at varying conditions using Protean II xi cells (Bio-Rad) (Table 1).
Silver staining of the gels was carried out after electrophoresis using a method described by Byun et al (2009) [12]. Homozygous and heterozygous patterns were identified from SSCP banding patterns.
Data analysis
IBM SPSS Statistics (Version 23, IBM, NY, USA) was used to perform the data analysis. Variant and genotypic frequencies were calculated using the Pop Gene 3.2 software. The variant and genotypic frequencies were tested for the deviation from the Hardy-Weinberg equilibrium (HWE) using Chi-squared analysis (Table 2). DNAMAN 5.2.10 Lynn on Bio soft was used to construct the phylogenetic trees. All p values at P < 0.05 were considered statistically significant.
Results and Discussion
SSCP patterns
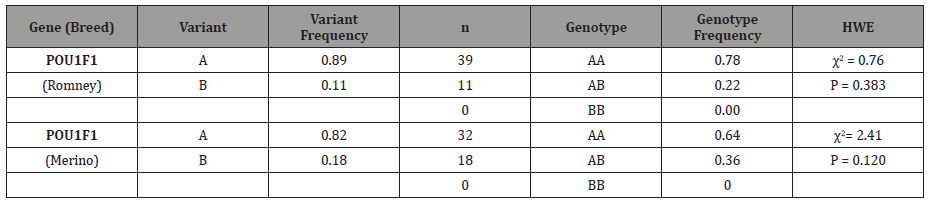
The exon 1 of POU1F1 exhibited limited heterozygosity with variants, AA and AB (Table 2). Fifty samples were investigated for the Romney breed, and another 50 for the Merino breed. For the two both sheep breeds, the major variant was the A variant, with a frequency of 89% (Romney) and 82% (Merino), while the B variant had a smaller frequency of 11% (Romney) and 18% (Merino) respectively. The variant frequencies did not deviate from the Hardy-Weinberg equilibrium.
Table 2: Variant and genotype frequencies for POU1F1 variants in NZ sheep breeds.
POU1F1 belongs to the POU-domain family of transcription factors which share a common N-terminal transactivation domain (TAD), and a C-terminal POU-specific domain (POUSD) and POU-homeodomain (POUHD). The TAD domain is crucial for the transactivation of target genes-GH, TSH and PRL, while high-affinity DNA binding is ensured by the POUSD and POUHD regions[13]. Previous research has linked polymorphisms in the exon 6 of POU1F1 with milk yield, and milk fat and lactose content [14]. Another study found association between POU1F1 and wool weight and fiber diameter [15].
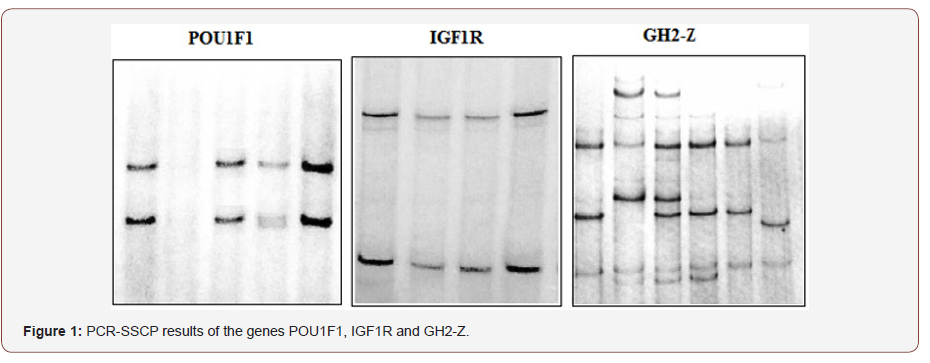
Exon 15 of IGF1R was homozygous for both Romney and Merino breeds, with no variations detected (Figure 1). This result could be attributed to the small sample size or could be due to the highly conserved nature of IGF1R owing to its function as an important housekeeping gene [16]. IGF1R plays an important role in growth, anabolism and homeostasis in mammals. Genes that are critical to the biological function of an organism are usually highly conserved in certain regions [17]. In contrast to this study, significant association has been reported between a polymorphism in IGF1R and body weight and average daily gain of coarse-wool ewes [18].
Multiple non-specific SSCP patterns were observed for GH2-Z (Figure 1). This is likely due to the gene duplication observed in ovine growth hormone. Many similar regions exist between GH1, GH2-N and GH2-Z.Combined GH2-N and GH2-Z variations have been demonstrated to bring about an approximate increase in milk production by 25% [19].
Allele and genotypic diversity
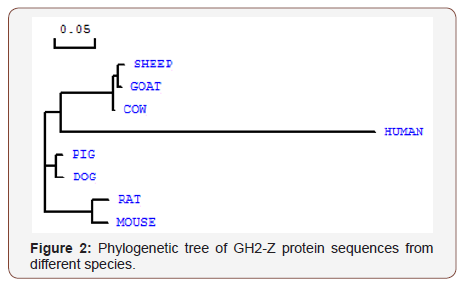
The evolutionary relationship between different species varies for different genes. For growth hormone, the genetic similarity between small ruminants, sheep and goat, are much closer than the link with the other mammalian breeds (Figure 2).
The limited diversity observed for POU1F1 in this study comes as no surprise as previous reports have shown coding region of ovine POU1F1 to be highly conserved as evidenced by its similarity with the bovine (98.2%), human (91.2%) and murine (86.2%) counterparts [20].
[The tree was constructed using the DNAMAN 5.2.10 Lynn on Bio soft package, based on NCBI accession numbers: Human, Homo sapiens (AY890602); Sheep, Ovisaries (DQ461615); Goat, Capra hircus (GU355687); Cow, Bos Taurus (KP221576); Pig, Sus scrofa (AY536527); Dog, Canisfamiliaris (AF069071); Rat, Rattus norvegicus (U62779) and Mouse, Mus musculus (NM_008117.3)]
Traits such as growth rate, meat and milk quality, and wool quality, dictate consumer choices and preferences. The animal breeding industry has undergone tremendous transformations in recent years, with marker-assisted selection superseding traditional methods of ‘visual appraisal’ [9]. This study provides insight into the nature of variability of three genes across two New Zealand sheep breeds. The GH2-Z gene was highly variable, POU1F1 was slightly variable, and IGF1R displayed no variability. Further work that could be done in this area with a larger sample size, as well as performing nucleotide sequencing of the variant patterns to ascertain the precise nature of the variation. Association studies can also be carried out to test possible links between variants and phenotypic traits.
Conclusion
Several chromosomes in sheep have been identified as QTLs for production traits such as growth and carcass traits, and milk production. Present study investigated three candidate genes found within these QTLs and their variant frequencies. The number of polymorphisms detected differed across the three genes studied, with GH2-Z having the most polymorphisms and IGF1R displaying no variations. Further research needs to be conducted into validating these variations and determining their associations with various production traits.
Acknowledgement
None.
Conflict of Interest
No Conflict of Interest.
References
- Campbell GS (1997) Growth-hormone signal transduction. The Journal of pediatrics 131(1): S42-S44.
- Vacca G, Dettori ML, Balia F, Luridiana S, Mura MC, et al. (2013) Sequence polymorphisms at the growth hormone GH1/GH2-N and GH2-Z gene copies and their relationship with dairy traits in domestic sheep (Ovis Aries). Molecular biology reports 40(9): 5285-5294.
- Gootwine E, Sise J, Penty J, Montgomery G (1993) The duplicated gene copy of the ovine growth hormone gene contains a PvuII polymorphism in the second intron. Animal genetics 24(4): 319-321.
- Rosenfeld RG (2006) Molecular mechanisms of IGF-I deficiency. Hormone Research in Pediatrics 65(1): 15-20.
- De la Rosa Reyna X, Montoya HM, Castrellon VV, Rincon AM, Bracamonte MP, et al. (2010) Polymorphisms in the IGF1 gene and their effect on growth traits in Mexican beef cattle. Genet Mol Res 9(2): 875-883.
- Chia DJ (2014) Minireview: Mechanisms of growth hormone-mediated gene regulation. Molecular Endocrinology 28(7): 1012-1025.
- Woollard J, Tuggle C,De Leon FP (2000) Rapid communication: localization of POU1F1 to bovine, ovine, and caprine 1q21-22. Journal of animal science 78(1): 242-243.
- Cohen LE, Wondisford FE, Radovick S (1996) Role of Pit-1 in the gene expression of growth hormone, prolactin, and thyrotropin. Endocrinology and metabolism clinics of North America 25(3): 523-540.
- Raadsma HW, Thomson PC, Zenger KR, Cavanagh C, Lam MK, et al. (2009) Mapping quantitative trait loci (QTL) in sheep. I. A new male framework linkage map and QTL for growth rate and body weight. Genetics Selection Evolution 41: 34.
- (2016) MIA. Export trends in selected markets up to March 2016. (Meat Industry Association of New Zealand (Inc)).
- Zhou H, Hickford J, Fang Q (2006) A two-step procedure for extracting genomic DNA from dried blood spots on filter paper for polymerase chain reaction amplification. Analytical Biochemistry 354(1): 159-161.
- Byun S, Fang Q, Zhou H, Hickford J (2009) An effective method for silver-staining DNA in large numbers of polyacrylamide gels. Analytical Biochemistry 385(1): 174-175.
- Bastos E, Avila S, Cravador A, Renaville R, Guedes Pinto H, et al. (2006) Identification and characterization of four splicing variants of ovine POU1F1 gene. Gene 382: 12-19.
- Ozmen O, Kul S, Unal EO (2014) Polymorphism of sheep POU1F1 gene exon 6 and 3’UTR region and their association with milk production traits. Iranian journal of veterinary research 15(4): 331.
- Negahdary M, Majdi S, Hajihosseinlo A (2014) Genetic Effect of IGF1, PIT1 and Leptin Genes on Wool Weights in Makooei Sheep. Electronic Journal of Biology 10: 46-51.
- Lei M, Peng X, Zhou M, Luo C, Nie Q, et al. (2008) Polymorphisms of the IGF1R gene and their genetic effects on chicken early growth and carcass traits. BMC genetics 9: 70.
- Barbieri M, Bonafe M, Franceschi C, Paolisso G (2003) Insulin/IGF-Isignaling pathway: an evolutionarily conserved mechanism of longevity from yeast to humans. American Journal of Physiology-Endocrinology and Metabolism 285(5): E1064-E1071.
- Proskura WS, Szewczuk M (2014) The polymorphism in the IGF1R gene is associated with body weight and average daily weight gain in Pomeranian coarsewool ewes. Pak Vet J 34: 514-517.
- Marques Md R, Santos IC, Carolino N, Belo CC, Renaville R, et al. (2006) Effects of genetic polymorphisms at the growth hormone gene on milk yield in Serra da Estrela sheep. Journal of dairy research 73(4): 394-405.
- Bastos E, Santos I, Parmentier I, Castrillo JL, Cravador A, et al. (2006) Ovis Aries POU1F1 gene: cloning, characterization and polymorphism analysis. Genetica 126(3): 303-314.
-
Ugonna J Ekegbu, Ishaku L Haruna, Ghassan Mahmoud, Huitong Zhou, Jon GH Hickford. Genetic Polymorphisms in New Zealand Sheep Breeds. World J Agri & Soil Sci. 1(2): 2018. WJASS.MS.ID.000506.
-
Marker-assisted selection, Polymorphism, Sheep, Gene, Animals and data collection, SSCP Patterns, Allele and genotypic diversity, Sheep breeds, Heterozygosity, Electrophoresis, Oligonucleotide primers, Lambs, Quantitative trait loci, Romney sheep
-

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.






