 Editorial
Editorial
Al Mana Group and Ucla Research Project for Molecular Genetic Studies of Degenerative Disorders of Saudi Children
Ahmed A Bashir* and Stanley F Nelson/p>
Department of Pediatrics, Al Mana General Hospital Jubail Saudi Arabia, Saudi Arabia
Ahmed A Bashir, Department of Pediatrics, Al Mana General Hospital Jubail Saudi Arabia.
Received Date: September 15, 2018; Published Date: April 30, 2019
Introduction
Dr. Bashir and his group at Al Mana Group frequently take care of and try to provide a genetic diagnosis for children with seriously disabling neurodevelopment and neuromuscular disorders. Even with comprehensive genetic testing, a clear molecular basis can remain elusive.Since many of these children are likely to have an undetected cause of their disorder, we propose here to develop an international collaboration between Dr. Bashir and Dr. Nelson to collect appropriate biomaterials for identification of causal mutations and push the envelope of detection of causal mutations by integrating RNAsequencing with whole genome sequencing and developing inexpensive models for improving the utility of RNAsequencing for neurodevelopmental disorders.Developmental disorders are frequently due to genetic mutations and can be detected in about 30% of cases using whole exome sequencing. Thus, the first screen for the included subjects is exomesequencing of an index case in the family. If a successful identification of a mutation is identified, these results are returned to the Al Mana Group to request ordering of a specifically developed molecular test through the UCLAOrphan Disease Testing Center on all relevant family members. This effectively serves as a powerful and thorough screen to identify those individuals who have a readily identifiable genetic disease.The remaining 70% of the cohort to be collected, will be subjected to short read based whole genome sequencing at UCLA. These data are able to sensitively detect, genome-wide, all SNVs and small indels (an average of 5 million per genome). We are thus well powered to observe DE NOVO mutations. However, there are over 60 de novo mutations per genome and only one is even potentially causal for a given rare autosomal dominant disease. Thus, interpretation of the functional consequence on RNA reading frame is imperative. Here we propose to use RNAsequencing of non-neural tissues to delineate ORF defects and intersect with the WGS DNA sequencing. We intend that this will result in higher rate of patient diagnosis at a molecular level and lead to insights of disease pathogenesis.For this project, we request consent and samples from Patients of Al ManaGroup which may include blood, skin, and saliva to study both RNA and DNA. We request fibroblast from skin punch in order to grow cells that can be de differentiated into iPSCs and then into a neural phenotype to induce expression of mRNAs expressed in brain or muscle. This is not a treatment study but may lead to insights towards new therapies.
Genome-wide diagnostic toolshaverapidly become an indispensable aspect of care for individuals with rare genetic disorders, frequently providing information on inheritance and prognosis, and in some cases direct therapeutic insights. The success of wholeexome sequencing (WES) in our clinical practice over the last 5 years strongly supports the proposition that largescale genetic information will be a major component of Precision Health throughout the lifespan[1]. However, even with the genetic definition of over 5,000 rare diseases, which in aggregate affect millions of Americans, a specific molecular diagnosis is returned after WES to only 30% of individuals tested[1-4]. While this is a major advance, 70% remain undiagnosed even with strong evidence of a genetic disorder. This is likely due to many factors, including incomplete genome coverage, poor detection of certain classes of structural variation (SV), especially within repeats, andthe vast identification of single nucleotide variants and SVs whose significance is uncertain.
Here, we propose to improve the detection of disease causing rare recessive and de novo autosomal dominant variants and improve the utility of whole genome sequencing (WGS)for genetic diagnosis of rare diseases.We propose to improve WGS coverage to better assess repeat regions, deletion/duplications, and SV using long-read DNA sequencing. Further, we propose to determine functional consequence through integrated transcriptome analysis of both high depth short-read and long-read RNA sequencing. Investigation of a DNA alteration that affects splicing or RNA abundance has long been recognized to be important (A,B,C) and has been demonstrated to substantially improve the interpretation of VUS[5]. However, clinical cDNA sequencing is not used broadly for diagnostics, andRNA-sequin this context has been largely limited to research, so theestablishment of clinical WGS/RNAseqtools developed here, developsamore complete genomics and bioinformatics infrastructure for comprehensive genomic analysis that is critical to future precision patient management more broadly.By conducting this project in well-selected patients with undiagnosed, and likely genetic, diseases, in whom WES has been negative, we leverage the tremendous ongoing clinical effort for rare disease diagnosis based on WES. All of the tools developed here are designed to be readily extended to all undiagnosed diseases, establishing a feasible framework for adoption by other clinical laboratories. Here, we pilot an integrated approach toinclude transcript tome data in the more routine interpretation of WGS of individuals with neurodevelopmental disorders(ND) by analyzing the transcriptome from patient derived blood and skin fibroblasts. To increase the likelihood of observing expression, these libraries are normalized, which allows us to observe about 85% of all neurological disease genes from one of these two non-neurological tissue sources. While these tissue types are not ideal, for ND the affected tissue (i.e. brain) is not readily possible to biopsy, thus alternate cell types are beneficial for understanding DNA mutation effects on mRNA, as they represent a broad sampling of gene expression and can be readily and ethically accessed in genetics clinics throughout our network, including at the County facilities. This lessens burden of participation and broadens accessibility, which are key aspects to implementation in relation to precision medicine. We hypothesize that implementing RNA-seqwhen jointly analyzed with more complete WGS that includes sensitive search for structural variants (SV)will substantially increase diagnostic rate. Identifying the genetic cause of an individual’s disease has immediate impact on their clinical care, providing clarity on inheritance within the family and guidance for treatment. The final WGS/RNA-seq clinical test will be complex but will in this proposal be implemented as a series of transferable protocols. The data analysis pipelines proposed here require substantial optimization on custom compute server configurations that include field programmable gated arrays(FPGA) through a collaboration with Falcon Computing, a commercial partner. This hardware acceleration has already reduced the server footprint by a factor of 10 and accelerated the task of WGS analysis of a human genome by 10-fold.
Project Plan
This demonstration project is intended todevelop and implement a novel and complex WGS/RNA-seq diagnostic for those individuals for whom even exomesequencing fails to identify the causal mutation.Here, we select for comprehensivegenomic analysis a core group of 50 patients and relevant family members seen in clinical practice by Dr. Bashir and colleagues in the Al Mana Group (total ~150 subjects)with pediatric neurologic or neuromuscular disorders. All individuals will be consented for genomic sequencing and provision of appropriate biomaterials.All 50 affected individuals will be sequenced by whole exomeseuqencing to screen for known disease-causing mutations.Most of these 50 individuals, from our experience of sequncing over 4,000 individuals at UCLA will be negative for a causal mutation. We estimate that 33 families will then be tested in the below molecular assays (approx. 100 individuals). This will provide the opportunity for new gene discovery and create the framework for ongoing collaboration between UCLA and Al Mana Group.In all cases, neither parent is affected, and the inheritance pattern is consistent with either a rare recessive model or a de novoautosomal dominant model. Strategically, ten of the selected individuals have an exome-identifiedVUS that has some evidence that the variantimpactsthe Open Reading Frame (ORF), and these along with variants in canonical splice acceptor/ donor sites will serve as positive controls of the RNA analysis to resolve mRNA effect of the DNA variant. Samples will be selected from the clinical judgment of Al Mana Group physicians, so that the breadth and utility of the approach is demonstrated across the group of relevant patients in Saudi Arabia.This workflow greatly benefits from the tremendous ongoing clinical efforts at our UC Health campuses to identify the genetic cause of rare diseases.We will observe the relative value of each WGS/RNA-seq technology in identifying mutations that disrupt ORFs of mRNA by comparing groups of patient TRIOs. Here, we will assay33TRIOs withaugmented short read 30x oversampling of the human genome using Illumina technology with comprehensive long read DNA sequencing to more sensitively observe SVs and deletions/duplications and determine their pathologic consequence. These data will be integrated with deep whole transcriptome RNA sequencing from tissue libraries using short read technologies.A subset of unsolved cases will have long read mRNA analysis to attempt resolve RNA consequence. The integration of these assays should greatly improve structural variation detection over the broadest possible size range and directly observe its consequence on mRNA abundance or ORF to augment interpretation.
Aim 1: Generation of Data.1a) Extend WgsData to Improve Identification of Structural Variants
Here we push the envelope of SV detection to augment the high sensitivity and accuracy of SNV and small INDEL detection from Illumina short read technology. To improve haplotyping, which simplifies and improves base calling, libraries for sequencing at HLI will be prepared using 10X GenomicsChromium Genome Library kit[6]. The major improvement will be in the arena of SV, which is not sensitively resolved by short reads: deletions and duplications in the range of a few hundred base pairs to kilobases are difficult to reliably detect genome-wide. Potential common sources of mutation that are invisible include tandem repeat array variations, inversions, deletions or duplications that are flanked by repeat sequences, and ALU and LINE insertions. Long repeats of high sequence similarity confound the ability to fully read a human diploid genome with short reads. Long single molecule reads greatly facilitate diploid human genome assembly when combined with short reads[7-9], providing unprecedented power to observe important mutation types such as (Figure1).
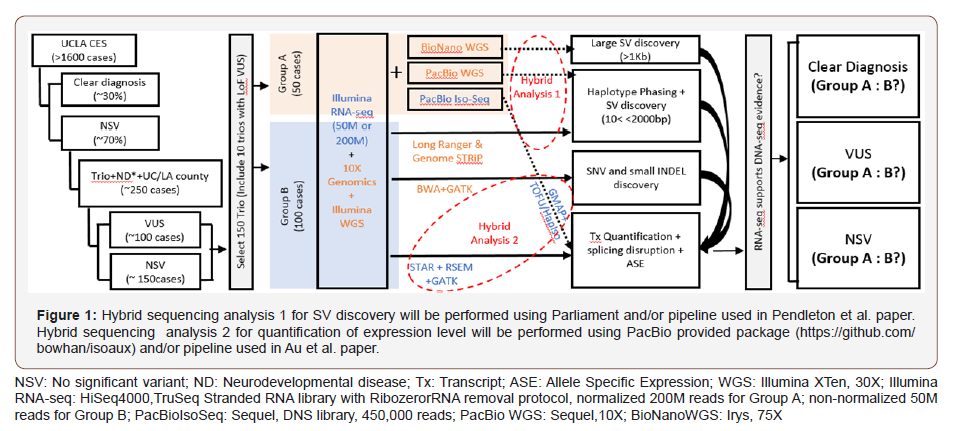
LINE insertions[10] or complex structural variation.The approach here is to have a set of complementary technologies to resolve a wide range of disease-causing mutations, ranging from single base variants through large SV. All 50 trios will be subjected to very long read single molecule BioNano GenomicsIrys assay in order to identify larger scale structural variation and to resolve complex, repeat laden genomic regions.Irys uses a two different restriction sequences to nick and tag the genome with a fluorescent marker at positions of known sequence, then through nano channels[11] measure the distance between marker which in effect makes a physical or optical map of the human genome[12-16], permitting detection of deletions as small as 1kb. Since the mean read length is 350kb, this technology allows spanning of complex and highly repetitive sequence in the human genome, and sizing of large tandem repeats. We have applied the Irys system to accurately detect large or complex deletions and partial gene duplications in DMDin both affected maleDMD patients and carrier mothers. Based on these experiences and technical capability, we will generate mean 75x genome coverage. There is about 5% failure to label at any given potential nicking site, and base level resolution is not obtained. However, at 75x mean coverage there is negligible contribution to miscalling of deletions/duplications in the range of 10kb to 300kb. Of known deletions in a pilot set of genomes, in the range of 5-50kb, we have 95% sensitivity of detection using two enzymes.Determining specificity and appropriate determination of support from short reads is an aim of the analytical plan.All SV will be mapped to predict consequence on mRNA abundance/ORF of all known ND genes genome-wide. Support for these mRNA effect predictions will be sought through RNA-seqand matched to the known disease model.
Aim 2: Augment DNA Sequencing with Normalized Transcriptome Analysis
The thorough DNA sequence generated in aim 1 will provide high-resolution SNVand SV information, but it will remain critical to determine consequence of variants on the ORF of mRNA through transcriptome analysis as many detected DNA variants will not be possible to interpret. As demonstrated recently, the addition ofRNAseq can greatly facilitate the determination of pathogenicity of seemingly benign variation detected by WES or WGS. For instance, missense or synonymous DNA variants detected by WEScan create strong splice acceptors or donors and result in aberrant splicing[5]. Similarly, deep intronic variants not within the canonical splice acceptor or donor sequences can create strong pseudo exons that are included in the mature mRNA and disrupt reading frame[5]. To increase sensitivity and to gain understanding of the potential to increase diagnostic yield from WGS, we will augment DNA sequence with RNA analysis[17,18].For neurodevelopmental disorders, the affected tissue type isoften not available (i.e. brain). Thus, use of more available tissue types, blood and dermal fibroblasts, still provides substantial information and allows the approach to be practically implemented within most genetics clinics, as blood draw and skin punch are routinely performed in clinics. It is obvious that since these cell types are not necessarily affected by the mutations causing brain disorders or other syndromic disorders, the affected gene may not be well expressed in these tissues and this does diminish the utility of RNA analysis. However, we note that without normalization, 50% of all neurodevelopmental disease genes are observed at greater than 2 FPKM in one of these cell types (internal reference data). To improve the depth of coverage of low expression genes, we will employ cDNA normalization strategy. Whole blood will be collected in PAXgene tubes and placed on ice for transport so that high quality long RNA can be purified safely within 24 hours. Skin punches will be placed in culture to generate mRNA from fibroblasts in 2-3 weeks. These two tissue types are nicely complementary in expression pattern[5]. To increase the proportion of the neurodevelopmentally-relevant transcriptome that are sufficiently covered to observe allele specific expression, we will subject the long fragment cDNA library to duplex specific nuclease (DSN) based normalization (Evrogen)[19-24]. All of the individuals in Group A that are not clearly resolved and fully interpretable from Aim 1, will be subjected to this normalized RNA analysis. Initial RNA Analysis will consist of 200 million fragments paired end 100bp reads on the Illumina platform, which allows observation of exon junctions from split read and read-pair mapping. Apparently causal defects will be confirmed by RT-PCR.
Aim 3:Develop and Implement Transferrable, State of the Art Tools for Efficient and Cost Effective Wgs Analysis and Integration with Rna-Seq Data
A)We will implement a state of the art pipeline for WGS analysis using the customized computing technology developed by Falcon Computing to accelerate analysis by an order of magnitude with field programmable gate arrays (FPGAs). This technology was originally developed by the Center for Domain Specific Computing (CDSC) at UCLAdirected by Prof. J. Congwith the support of NSF and Intel under the Expeditions in Computing and Innovation Transition (InTrans) Programs. A single Falcon computing appliance, with a physical footprint of 2Userver, can perform whole genome alignment and variant calling through GATK best practice pipeline in an average of 5 hours (100+ hours in our high RAM compute nodes). This acceleration removes a key obstacle to the use of WGS interpretation in the clinical setting and substantially lowers barriers towards adoption. Under the direction of Prof. Cong with Prof. E. Eskin, these appliances will be tuned to accelerate alignments and integration of the 75x BioNano genomics data Reducing compute times is an important to implement in clinical practice allowing greater flexibility to meet reasonable turnaround time and creating a more robust computational pipeline.
Aim 4: Assess Relative Benefit of Augmented Genomic Analyses
We will integrate the more complete WGS with RNA-seq, withtranscript abundance, splicing, phasing of variants in mRNA[25],and allele specific expression data to both guide interpretation of WGS and provide evidence for variant pathogenicity. We willassess the improvement in diagnostic rate and specificity of RNA-seq coupled with WGS, as compared with WES alone (E). We will specifically determine the rate of pathogenic or likely pathogenic mutation/variant identification for the 33individuals with completer WGS more coupled with RNA-seq relative to WES data, and compare with the inferences possible from WGS RNA-seqof whole blood from short read technology alone.We will further enumerate the number of loss-of-function DNA variants, based on the combined analysis withRNA and better SV resolution across all parental samples that are consistent with detection of a carrier state for any rare recessive disease as an additional measure of pathogenic variant detection assessment. Primarily, the genomic and transcriptomic data generated will be assessed for their utility in diagnosing children with rare ND disorders, but we will also broadly consent families for datasharingsuch that these data can be used as a reference. Since fibroblasts will be available as well additional genome-wide or targeted datasets can be gathered in the future to assess relevance to the identification of rare and common variation.We will also use these data to bench mark improvements in DNA sequence assembly algorithms.Thus, this pilot project serves not only to extend the sensitivity of genomic assessment in rare diseases,but will be a flagship project of central importance to the development of precision medicine on the UCLA campus and in partnership with the Al Mana Group to explore improvements in genetic diagnosis and utility of comprehensive genetic assessments made possible by genome technology.Because of this importance to our efforts at UCLA, we are able to leverage personnel and computational resources through the launch of the UCLA Precision Health Institute.These complementary activities enable a more robust diagnostic developmental pipeline.Our project builds specific collaborations with BioNano Genomics and Falcon Computing, HLI, and uses two California technologies (Illumina and Pacific Biosciences) and partners with all 5 UC campuses and some of the LA County facilities.By completing these aims we will have made major inroads in developing translational infrastructure for precision medicine for undiagnosed patients, and integrating RNA-seq and WGS into clinical practice, important steps towards developing best practices for Precision Health in California and in Saudi Arabia.We note that while the above pilot process is expensive, ongoing improvements in technology development, and perhaps a tiered diagnostic approachwill permit more thorough genome assessment at reasonable costs. For the participants of this research study, each predicted causal DNA mutation identified from this pilot will not be directly reported to the physician/ family, but rather after discussion in the GDB with the ordering physician,specific testing is ordered through the UCLA Orphan Disease Testing Center.All relevant and interested individuals within the family can be tested with the developed family-specific DNA diagnostic. A clinical report is generated through the UCLA Molecular Pathology Diagnostic Lab with Dr. Wayne Grody.
Precision Medicine Capabilities
This project will develop and assess a more comprehensivediagnostic process intended toimprove the full interpretation of individual human diploid genomes and reveal mRNA consequence of non-coding DNA variants with implementation into clinical practice. Our ability to improve our competency in clinical genome interpretation is an essential aspect of precision medicine. This collaboration will informtwoway communication channels between laboratory and clinicians. The developed enterprise will enhance novel gene discoveryfor ultra-rare diseases, which can lead to strong genetically tailored therapeutic insights.Finally, this pilot will allow us to develop efficient algorithms for the joint analysis of RNA and DNA and implement them on more optimized hardware solutions, which are transferable to other centers.
Impact for Patients
In the last 3 years, we and our colleagues have performed CESon over 1600 patients. Per the NIH, ~7% of patients asking for their help have an undiagnosed disease. The cost of these clinical investigations, in the absence of clear diagnostic protocols, is high. For example, the estimated lifetime medical care cost for a person with intellectual disability, often associated with undiagnosed disorders, was around $1M in 2003 (F) and contributes to the economic strains on medical systems. Improving diagnostic yield provides closure for families affected by rare diseases, improves the clarity of prognostic information, and allows for meaningful genetic counseling regarding family planning[26,27].
Anticipated Challenges and Proposed Solutions
An integrated WGS/RNA-seq diagnosticrequires several separate analytical inputs, tracking, and computational integration. Critical bottlenecks that slow the proposed diagnostic test will be systematically addressed. We will be continuously altering software and hardware components to harmonize timing of component analyses.Interpretation of the final data will be a challenge, even with interpretation through the GDB. Even long read technologies such as Iso-Seq do not read through long transcripts such as DMD (~14Kb) or TTN (~110Kb), and allele specific expression of low expression genes may not be adequately observed from whole blood or fibroblasts. Thus, gaps will remain. Development of individual induced neural cells using iPSCs and more sequencing of diverse populations canreduce these gaps. The participants arehighly motivated families who have already undergone CES, but our physicians may experience difficulty enrolling due to patient movement within the health system or interest in data sharing. Since there is a growing group of exome-negative individuals, we are confident of recruitment goals.Establishing a full CAP/CLIA system is the goal at the end of the project, and it will be challenging to codevelop standard operating procedures and implement within the Molecular Diagnostics Laboratories as all aspects of the technical system, analytical system and interpretation system need to be fully validated to permit clinical reporting of variants. We may need to rely on ODTC reporting of identified variants. We faced similar challenges during the initial 6 month implementation of CES,among the first in the US. Given the clinical, bioinformatic, hardware, and genomic expertise assembled for this project, we anticipate similar progress.
Project Team
Dr. S. Nelsonhas 25 years of genomics experience and established CES for rare disease diagnosis on the UCLA campus will serve as overall PI on the site campus with additional key leadership from Dr. A. Bashir of the Al Mana Group. Dr. Bashir will effectively serve the role as physician and consenter of the relevant patients in Saudi Arabia, and as postdoctoral fellow at UCLA in the Nelson Lab which will allow ?Dr. Bashir to participate in all aspects of the assay development, interpretation, and drafting of manuscripts from the collaborative work.Much of the work will occur under the auspices of the Department of Human Genetics at UCLA and the Institute for Precision Health at UCLA.
Acknowledgement
None.
Conflict of Interest
No Conflict of Interest.
References
- Hane Lee, Joshua L Deignan, Naghmeh Dorrani, Samuel P Strom, Sibel Kantarci, et al. (2014) Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA 312(18): 1880-1887.
- Arboleda VA, Lee H, Dorrani N, Zadeh N, Willis M, et al. (2015) De novo nonsense mutations in KAT6A, a lysine acetyl-transferase gene, cause a syndrome including microcephaly and global developmental delay. Am J Hum Genet 96(3): 498-506.
- Yang, Y, Donna M Muzny, Fan Xia, Zhiyv Niu, Richard Person, et al. (2014) Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 312(18): 1870-1879.
- Fogel BL, Lee H, Deignan JL, Strom SP, Kantarci S, et al. (2014) Exome sequencing in the clinical diagnosis of sporadic or familial cerebellar ataxia. JAMA Neurol 71(10): 1237-1246.
- Cummings BB, Marshall JL, Tukiainen T, Lek M, Donkervoort S, et al. (2017) Improving genetic diagnosis in Mendelian disease with transcriptome sequencing. Bio Rxiv.
- Kitzman JO (2016) Haplotypes drop by drop. Nat Biotechnol 34(3): 296- 298.
- Mostovoy Y, Levy-Sakin M, Lam J, Lam ET, Hastie AR, et al. (2016) A hybrid approach for de novo human genome sequence assembly and phasing. Nat Methods 13(7): 587-590.
- Pendleton M, Sebra R, Pang AW, Ummat A, Franzen O, et al. (2015) Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat Methods 12(8): 780-786.
- Shi L, Guo Y, Dong C, Huddleston J, Yang H, et al. (2016) Long-read sequencing and de novo assembly of a Chinese genome. Nat Commun 7: 12065.
- Hancks, DC, Kazazian HH (2012) Active human retrotransposons: variation and disease. Curr Opin Genet Dev22(3): 191-203.
- Cao H et al. (2002) Fabrication of 10 nm enclosed nanofluidic channels. Applied Physics Letters 81: 174-176.
- Lam ET, Hastie A, Lin C, Ehrlich D, Das SK, et al. (2012) Genome mapping on nanochannel arrays for structural variation analysis and sequence assembly. Nat Biotechnol30 (8): 771-776.
- Roberts RJ, Carneiro MO, Schatz MC (2013) The advantages of SMRT sequencing. Genome Biology 14(7): 1-4.
- Lee, H. et al. (2014) Error correction and assembly complexity of single molecule sequencing reads. bioRxiv.
- Koren, S. et al. (2012) Hybrid error correction and de novo assembly of single-molecule sequencing reads. Nat Biotechnol 30.
- Mangul, S, Harry Yang, Farhad Hormozdiari, Elizabeth Tseng, Eleazar Eskin, et al. (2016) HapIso: An Accurate Method for the Haplotype- Specific Isoforms Reconstruction from Long Single-Molecule Reads. bioRxiv pp 80-92.
- Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ (2008) Deep surveying of
alternative splicing complexity in the human transcriptome by highthroughput
sequencing. Nat Genet 40(12): 1413-1415.
- Chi, K.R. (2016) Finding function in mystery transcripts. Nature 529: 423-425.
- Zhulidov PA, Bogdanova EA, Shcheglov AS, Shagina IA, Vagner LL, et al. (2005) A method for the preparation of normalized cDNA libraries enriched with full-length sequences. Bioorg Khim 31(2): 186-194.
- Tilgner H, Grubert F, Sharon D, Snyder MP (2014) Defining a personal, allele-specific, and single-molecule long-read transcriptome. Proc Natl Acad Sci U S A 111(27): 9869-9874.
- Vreeswijk MP, van der Klift HM (2012) Analysis and interpretation of RNA splicing alterations in genes involved in genetic disorders. Methods Mol Biol 867: 49-63.
- Gaildrat P, Killian A, Martins A, Tournier I, Frébourg T, et al. (2010) Use of splicing reporter minigene assay to evaluate the effect on splicing of unclassified genetic variants. Methods Mol Biol 653: 249-257.
- Walker LC, Whiley PJ, Houdayer C, Hansen TV, Vega A, et al. (2013) Evaluation of a 5-tier scheme proposed for classification of sequence variants using bioinformatic and splicing assay data: inter-reviewer variability and promotion of minimum reporting guidelines. Hum Mutat 34(10): 1424-1431.
- Castel SE, Mohammad, P, Chung WK, Shen Y, Lappalainen T (2016) Rare variant phasing and haplotypic expression from RNA sequencing with phASER. Nat Commun 7: 12817.
- McCullough LB, Slashinski MJ, McGuire AL, Street RL, Eng CM, et al. (2016) Is Whole-Exome Sequencing an Ethically Disruptive Technology? Perspectives of Pediatric Oncologists and Parents of Pediatric Patients With Solid Tumors. Pediatr Blood Cancer 63(3): 511-515.
- Fogel BL, Satya-Murt, S, Cohen BH (2016) Clinical exome sequencing in neurologic disease. Neurol Clin Pract 6(2): 164-176.
-
Ahmed A Bashir, Stanley F Nelson. Al Mana Group and Ucla Research Project for Molecular Genetic Studies of Degenerative Disorders of Saudi Children. Glob J of Ped & Neonatol Car. 1(1): 2019. GJPNC.MS.ID.000505.
-
Degenerative disorders, Genetic diagnosis, Neurodevelopmental disorders, Neuromuscular disorders, RNA Sequencing, Genetic mutations, UCLA Orphan disease, Autosomal dominant disease, Diagnosis, Pathogenesis, Blood, Skin, DNA, Exome sequencing, cDNA, mRNA, Comprehensive genomic analysis, Skin fibroblasts, Blood fibroblasts, Neurological disease, Non-Neurological tissue
-

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
- Abstract
- Introduction
- Project Plan
- Aim 1: Generation of Data.1a) Extend WgsData to Improve Identification of Structural Variants
- Aim 2: Augment DNA Sequencing with Normalized Transcriptome Analysis
- Aim 3:Develop and Implement Transferrable, State of the Art Tools for Efficient and Cost Effective Wgs Analysis and Integration with Rna-Seq Data
- Aim 4: Assess Relative Benefit of Augmented Genomic Analyses
- Precision Medicine Capabilities
- Impact for Patients
- Anticipated Challenges and Proposed Solutions
- Project Team
- Acknowledgement
- Conflict of Interest
- References






