 Review Article
Review Article
Analytical Framework for the Mathematical Modelling and Analysis of COVID-19 Transmission Dynamics in Latent and Active Stages
F C KIWU-LAWRENCE1*, F N NWOBI2, E U OHAEGBULEM2
1Department of Statistics, Abia State University Uturu, PMB 2000, Uturu, Abia State, Nigeria
2Department of Statistics, Imo State University Owerri, PMB 2000, Owerri, Imo State, Nigeria
F C KIWU-LAWRENCE, Department of Statistics, Abia State University Uturu, PMB 2000, Uturu, Abia State, Nigeria
Received Date: December 15, 2025; Published Date: February 12, 2026
Abstract
In this paper, an analytical framework of the mathematical model for assessing the epidemic dynamics of the latent and active stages of COVID-19 disease is proposed. The population group was divided into six categories. The biological feasibility of the developed mathematical model was studied by verifying the properties such as the existence, boundedness and the non-negativity of the solutions. The analytical behaviour of the disease-free equilibrium point, the basic reproduction numbers, and the local stability analysis for the latent and active stages of COVID-19 were studied using the linearization approach. The results show that the solutions of the system outcomes are positive and finite within the region, with the disease-free equilibrium point, the population size is stable. The basic reproduction number depends on two distinct transmission pathways: the latent and the active COVID-19 effective contact rates, suggesting that the COVID-19 latent (asymptomatic) stage contributes in the disease transmission patterns. This study demonstrates the importance of considering the latent stage of a disease in the modelling of a disease dynamics and its implications for epidemiological studies.
Keywords:COVID-19; latent; active; disease free equilibrium; basic reproduction number (R0); stability
Introduction
The COVID-19 pandemic started in Wuhan, China, in December 2019 [1]. This disease, caused by Severe Acute Respiratory Syndrome Corona Virus 2 (SARS-CoV-2), recorded in its first week of pandemic, asymptomatic cases of barely 1%. Among infected people, 81% had mild symptoms, 14% were severe, and 5% were critical [2]. Since its emergence, COVID-19 has spread globally, infecting over 676 million people, resulting in 6.8 million deaths, and promoting the administration of 13.3 billion vaccine doses, with economic losses amounting to trillions of USD [3,4]. The epidemic dynamics of COVID-19 are characterized by presymptomatic transmission events, suggesting a shorter latent period compared to the incubation period [5]. The latent period is typically estimated as the time between infection and detectable virus in a respiratory specimen. Common symptoms of SARS-CoV-2 infection include fever, cough, and fatigue, with some individuals experiencing stuffy or runny nose and diarrhoea. Severe cases can lead to acute respiratory distress syndrome, septic shock, and death [6]. As a highly contagious infectious disease, its sources of infection include both confirmed cases and also asymptomatic carriers [6,7].
Asymptomatic patients of SARS-CoV-2 infection are often without clinical symptoms, but test positive for the viral nucleic acid test. Most asymptomatic patients are found during the screening of close contacts. Because the laboratory tests are performed at an early stage, these asymptomatic cases go on to develop illness after screening. According to [8], the most common symptoms of COVID-19 include fever, cough, and fatigue, while Symptoms of severe COVID-19 disease are high temperature, loss of appetite, shortness of breath, persistent pain in the chest, etc. People of all ages who experience any of the aforementioned symptoms should seek medical care immediately, as some people may experience severe illness, which can lead to hospitalization and death. The primary mode of transmission of COVID-19 is through respiratory droplets generated during talking, coughing or sneezing by an infected person, which can land on surfaces or directly on another person’s mouth, nose, or eyes, facilitating infection.
Therefore, it is important to maintain physical distance, wear masks, and practice good hand hygiene to reduce the risk of transmission. Various models have been developed to explain COVID-19 dynamics, simulating strategies to mitigate the outbreak and reduce the peak in daily cases, hospital admissions, and fatalities. [9] described modelling COVID-19 spread in Germany, its assessment, and possible scenarios. They simulated different strategies for the mitigation of the current outbreak, to help in slowing down the spread of the virus and reducing the peak in daily diagnosed cases. Their results showed that a partial and gradual lifting of introduced control measures could be possible if accompanied by further increased testing activity, strict isolation of detected cases, and reduced contact with risk groups. [10] developed mathematical modelling of COVID-19 dynamics in Ukraine.
Therefore, it is important to maintain physical distance, wear masks, and practice good hand hygiene to reduce the risk of transmission. Various models have been developed to explain COVID-19 dynamics, simulating strategies to mitigate the outbreak and reduce the peak in daily cases, hospital admissions, and fatalities. [9] described modelling COVID-19 spread in Germany, its assessment, and possible scenarios. They simulated different strategies for the mitigation of the current outbreak, to help in slowing down the spread of the virus and reducing the peak in daily diagnosed cases. Their results showed that a partial and gradual lifting of introduced control measures could be possible if accompanied by further increased testing activity, strict isolation of detected cases, and reduced contact with risk groups. [10] developed mathematical modelling of COVID-19 dynamics in Ukraine.
Methodology
This section discusses the methods used in the COVID-19 SLAIIR model’s mathematical modeling. The model’s solutions include positivity and boundedness, as proposed in [13], which proves that the system’s solutions are positive and bounded. The integration factor method from [14] transforms the differential equations into an easily integrable form, ensuring biological meaningful results. The Disease-Free Equilibrium (DFE) point, is found by solving the right-hand side of the system of (4(a) − ( f )) to zero, representing a population state where COVID-19 is no longer present. This is typically characterized by zero infected individuals. The basic reproduction number, R0 , is a key parameter in epidemiological models, de ned as the number of secondary infections from a single primarily infected individual [15]. The latent and active basic reproduction numbers of the COVID-19 SLAIIR model was calculated using the Next Generation Matrix method (NGM) given by [16] as follows:
From (4(a) − ( f )) , we perform the following operations:
a. Regroup the equations in the model into disease and nondisease
classes.
b. Express disease class dynamics as  ,
where f includes new infections and v includes transitions
and removals.
,
where f includes new infections and v includes transitions
and removals.
c. Construct matrices F and V by partial derivatives of f
and v w.r.t. infected compartments at the disease- free equilibrium
(LC ,A C,ILC , and IAC respectively).
d. Calculate the next generation matrix K = FV−1 .
e. Obtain the eigenvalues of K = FV−1.
f. 0 R is the largest spectral radius of K,ρ (K) .
Finally, we examine the local stability near the disease-free equilibrium, to help us determine if small outbreaks die out or grow. We perform the local stability analysis of equilibrium points, as described by [13]. To achieve this, we use the linearization approach, given in [13], and perform the following operations;
a. Identify equilibrium points.
b. Linearize the system at the equilibrium point.
c. Compute the Jacobian matrix.
d. Evaluate eigenvalues: stability requires all eigenvalues to
have negative real parts.
Model Formulation
The SLAIIR mathematical model for the COVID-19 transmission
dynamics is formulated by dividing the population into six classes,
namely:
a. S (t ): Susceptible individuals who are at risk of
contracting COVID-19, they have not yet been infected and do not
have immunity to the disease;
b.LC (t ) : The latent COVID-19 infected individuals who just
contracted (got infected with) COVID-19, they represent individuals
in the incubation period who are asymptomatic;
c. A C(t ) : The active COVID-19 infected individuals who have
full blown COVID-19, they are symptomatic;
d. ILC(t ): Individuals infectious with latent COVID-19, who
have compromised immune system, are pre- symptomatic and can
transmit COVID-19;
e.IAC (t ) : Individuals infectious with active COVID-19, they
are experiencing symptoms of COVID-19, have high viral load,
making them a potential source of infection for others particularly
in close contact settings;
f. R(t ) : Individuals recovered from both latent and active
stages of COVID-19.
These classes represent various stages of COVID-19 infection, enabling the modeling of the disease spread. With these, the total population, N (t ) , is the sum of (a − f ) at time, t , given by:

Model Assumptions
For our proposed SLAIIR model, the following assumptions are
made:
a. The susceptible class increases by the recruitment of
people at a rate, Λ .
b. All individuals are subject to natural death, at rate, μ .
c. The transmission of disease occurs when susceptible
individuals come in contact with infected individuals, be it latent
COVID-19 or active COVID-19, by a force of infection, (the incidence
is of bilinear mass action). The forces of infections are given as:


where,
• 1 C β L is the effective contact rate for latent COVID-19; and
• 2 C β A is the effective contact rate for active COVID-19.
d. Individuals from the COVID-19 latent class, LC , become
infectious ( ILC) at a rate, v , or recover from latent COVID-19 at
a recovery rate, ϕ .
e. Individuals from the active COVID-19 class, AC , become
infectious (IAC ) at a rate, , or recover from active COVID-19 at a
recovery rate, π .
, or recover from active COVID-19 at a
recovery rate, π .
f. Infectious latent COVID-19, ILC and infectious active
COVID-19,
IAC , may recover at rates, ψ and γ , respectively, or
die due to COVID-19 disease-induced death rates of
δLC and δAC
, respectively.
The schematic diagram of the proposed (SLAIIR) model is illustrated in Figure 1. Thus, the governing mathematical model can be framed into the following system of equations:







where,
λLC and λAC are as given in (2) and (3), respectively; and with
initial conditions,

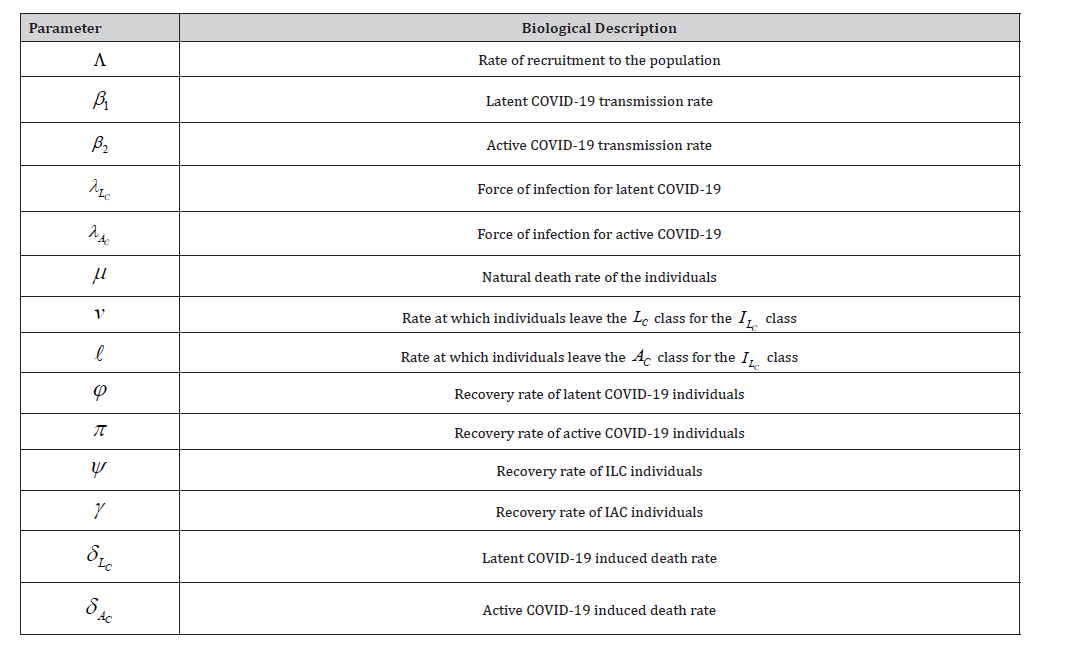
The descriptions of model parameters are presented in Table 1.
Table 1: The Description of the Proposed SLAIIR Model Parameters.
Model Analysis
Positivity and Boundedness of Solutions
This theorem ensures the model’s solutions are biologically realistic and acceptable to human populations, being both positive and bounded within a feasible range.
Theorem 3.1. All solutions of the SLAIIR model are positive and bounded in the region;

Proof. For a positive set Ω , and non-negative initial conditions (as given in (5)), each vector eld is demonstrated to remain non negative. Thus






The change in total population, given as: N(t)=S+ LC + AC + IL C + IAC + R , at time, t , is governed by;

Substituting each component of (6(a) − ( f )) into (7); and cancelling out, gives;

Equation (8a) can be re-written for convenience as;

Solving (8b) using the integration factor, we have:

Integrating both sides gives;

and then dividing both sides by eμt gives;

With the given the initial condition, 0 ≤ N (0) , as t →0 ,

Hence, the proof con rms that the solution of the SLAIIR model remains non-negative and bounded; ensuring biological meaningful results.
The Disease-Free Equilibrium Points (DFE)
Setting the disease states LC= AC = IL C = IAC = 0 in (4(a) − ( f )) , yields the disease-free equilibrium points, we have for LC:Λ−β1(0+0)S−β2(0+0 )S − μ S = 0 . Performing the same operations for C AC , IL C and IAC
we obtain the disease-free equilibrium points as:

The Basic Reproduction Number
From (4(a) − ( f )) , we have:


We obtain Jacobian matrices F and V by partial derivatives of f and v w.r.t. (LC ,AC ,IL C ,and IAC ) respectively, noting that for β1(LC+ IL C) S ,


Performing the same operations for other entries in f and v , we obtain;


After performing necessary matrix operations, the inverse of (18) is obtained as,

And the next generation matrix, FV −1 , as:

The eigenvalues (λ ) of FV −1 are:

and λ3 = λ4 with multiplicity 0 .
This follows that the basic reproduction numbers for the latent and active COVID-19 SLAIIR model are:

and

The overall basic reproduction number for the COVID-19 SLAIIR model is,

These results show that:
 depends on two distinct transmission pathways,
represented by β1
and β2.
depends on two distinct transmission pathways,
represented by β1
and β2.
 relates to the progression from latent COVID-19 to the
infectious latent COVID-19 compartment, denoted by v .
relates to the progression from latent COVID-19 to the
infectious latent COVID-19 compartment, denoted by v .
 relates to the progression from active COVID-19 to the
infectious active COVID-19 compartment, denoted by
relates to the progression from active COVID-19 to the
infectious active COVID-19 compartment, denoted by  .
.
The Local Stability Analysis
The Local Stability of steady state solutions is examined using linearization approach. The COVID-19 SLAIIR
model’s Jacobean matrix is assessed for stability evaluation at
 is:
is:

The eigenvalues are −μ of multiplicity 2. The other four are obtained from the reduced matrix;

From the 4× 4 sub matrix (25), the disease specific compartments can be obtained, that is, sub matrices corresponding to the latent and active stages of COVID-19.
The Submatrix corresponding to the latent COVID-19 is:

with

and

Eigenvalues for the sub-matrix are


According to the method of analysis of local stability, the trace is negative if

and the determinant is positive if

Hence, the DFE point,  , of the latent COVID-19 is
(a) Stable if
, of the latent COVID-19 is
(a) Stable if  <1
(b) Unstable if
<1
(b) Unstable if  >1
>1
Also, the sub-matrix corresponding to the active COVID-19 is

with

and

The trace is negative if

and the determinant is positive if

So, the DFE point,  , of the active COVID-19 is
(a) Stable if
, of the active COVID-19 is
(a) Stable if  <1, and
(b) Unstable if
<1, and
(b) Unstable if  >1.
>1.
Result Discussion
The global spread of COVID-19, caused by SARS-CoV-2, has led to substantial health and economic impacts. The pandemic, which can be asymptomatic (latent) or symptomatic, has rapidly spread to over 200 countries, continuing to inflict severe public health and socio-economic burden in many parts of the world. The SLAIIR compartmental epidemic model developed in this study, addresses the asymptomatic stage of the disease, among other stages. The positivity and boundedness of solutions showed that the solutions of the model remain positive for all time, indicating that the compartments are non-negative. The solutions are bounded, indicating that the total population is finite. These conditions imply that the epidemic will eventually die out. The DFE represents a state where the disease has been eradicated from the population. The DFE point shows that the susceptible compartment is at a steady state, indicating a constant population with no infection. All infected compartments are zero, indicating no infections in the population at the DFE. For our proposed SLAIIR model, our R0 is influenced by two distinct transmission pathways, indicating that COVID-19 can spread through multiple routes, (e.g., direct contact, airborne transmission, etc.). It suggests heterogeneity in transmission, where different routes contribute differently to the overall transmission dynamics. The local stability shows mixed stability, where the nonzero eigenvalues determine stability in some directions, while the zero eigenvalues indicate neutral stability in other directions. The results imply that the disease exhibits slow but impacting dynamics during latency, contrasting with faster dynamics during infection recovery.
Conclusion
This study introduces a new SLAIIR mathematical model to capture COVID-19 transmission dynamics, en- compassing latent and active phases. The model’s solutions are proven to be positive and bounded, ensuring a realistic depiction of disease spread. Also, equilibrium points were estimated and analyzed for stability in relation to the basic reproduction numbers of the latent and active stages of the disease. In both stages of the disease, we proved that the basic reproduction numbers depend on two distinct transmission pathways. The results of this analytical approach reveals that the disease-free equilibria are locally stable if R0 <1 , and unstable otherwise. This critical examination of the proposed SLAIIR mathematical model of the COVID-19 epidemic dynamics with latent and active stages sheds light on the role played by the latent stage of COVID-19 disease in the transmission dynamics of the disease.
References
- Zhu N, Zhang D, Wang W, Li X, Yang B, et al. (2020) 'A novel Coronavirus from patients with pneumonia in China. N Engl J Med 382(8): 727-733.
- Wu Z, McGoogan JM (2020) Characteristics of and Important Lessons from the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72314 Cases from the Chinese Center for Disease Control and Prevention. JAMA 323(13): 1239-1242.
- (2023) World Health Organization Coronavirus (COVID-19) Dashboard.
- Kolahchi Z, De Domenico M, Uddin LQ, Cauda V, Grossmann I, et al. (2021) COVID-19 and Its Global Economic Impact. Adv Exp Med Biol 1318: 825-837.
- Fine PE (2003) The interval between successive cases of an infectious disease. Am J Epidemiol 158(11): 1039-1047.
- He X, Lau EHY, Wu P, Deng X, Wang J, et al. (2020) Temporal dynamics in viral shedding and transmissibility of COVID-19. Nat Med 26(5): 672-675.
- Huang C, Wang Y, Li X, Ren L, Zhao J, et al. (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395(10223): 497-506.
- World Health Organization (2021) Coronavirus disease (COVID-19) pandemic.
- Barbarossa MV, Fuhrmann J, Meinke JH, Krieg S, Varma HV, et al. (2020) Modeling the spread of COVID-19 in Germany: Early assessment and possible scenarios. PLoS ONE 15(9): e0238559.
- Semenova Y, Pivina L, Khismetova Z, Auyezova A, Nurbakyt A, et al. (2020) Anticipating the Need for Healthcare Resources Following the Escalation of the COVID-19 Outbreak in the Republic of Kazakhstan. J Prev Med Pub Health 53(6): 387-396.
- Ramezani SB, Amirlati A, Rahimi S (2021) A novel compartmental model to capture the nonlinear trend of COVID-19. Comput Biol Med 134: 104421.
- Odagaki T (2023) New compartment model for COVID-19. Sci Rep 13(1): 5409.
- Brauer F, Castillo-Chavez C (2012) Mathematical models in population biology and epidemiology. Springer.
- Perko L (2001) Differential equations and dynamical systems (3rd ed.). Springer-Verlag.
- Mekonen K, Obsu L (2022) Mathematical modeling and analysis for the co-infection of COVID-19 and tuberculosis. Heliyon 8 (10): e11195.
- van den Driessche P, Watmough J (2002) Reproduction numbers and sub-threshold endemic equilibria for compartmental models of disease transmission. Math Biosci 180: 29-48.
-
F C KIWU-LAWRENCE*, F N NWOBI, E U OHAEGBULEM. Analytical Framework for the Mathematical Modelling and Analysis of COVID-19 Transmission Dynamics in Latent and Active Stages. Annal Biostat & Biomed Appli. 7(1): 2026. ABBA.MS.ID.000654.
COVID-19; latent; active; disease free equilibrium; basic reproduction number (R0); stability; iris publishers; iris publisher’s group
-

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.






