 Research Article
Research Article
Validation and Clinical Use of the Non-Invasive Prenatal Test “Veracity”
Valentina Gnetetskaya*, Ekaterina Kuznetsova, Yulia Tarasova and Mark Kurtser
Genetics Centre of Mother and Child Groups of Companies, Russia
Valentina Gnetetskaya, Head of Medical Genetics Centre of Mother and Child groups of companies, Prenatal Medical Centre, Moscow, Russia.
Received Date: April 27, 2020; Published Date: May 05, 2020
Introduction
Cell-free DNA has been utilized as a novel analyse for the development of non-invasive approaches to prenatal genetic testing using different methodologies [1-4]. Following the publication of a number of major research and clinical studies that revealed high accuracy to detect fetal aneuploidies, the use of noninvasive prenatal testing (NIPT) has been widely adopted in clinical practice. NIPT technologies provide significant improvements over conventional invasive prenatal testing and consequently, international bodies endorse NIPT as a routine screening option [5,6]. This has resulted in many institutions adopting NIPT within the scope of standard of care for autosomal and sex chromosomal aneuploidy detection (SCA). Mother and Child Clinic is a primary clinical centre certified by the Russian Ministry of Health and specialises in screening and diagnosing prenatal chromosomal aneuploidies. NIPT has been offered as a prenatal screening option in our clinic since (May 2017). This study aims to summarize the NIPT results and clinical performance of NIPT in the detection of trisomy 13,18,21 and SCAs using the Veracity non-invasive prenatal test in a cohort of 1382 samples among mixed-risk participants. In this report we convey a complete and robust clinical picture of the performance of Veracity under routine NIPT testing conditions and describe examples of unique clinical cases.
Materials and Methods
Patient cohort
This study included 1382 pregnant women between the ages 18-52 between 9-28 gestational weeks who visited Mother and Child Clinic from May 2017 until October 2018. Of these women 1325 were singleton pregnancies, 37 were twin pregnancies, 20 were vanished twin pregnancies and 169 conceived by in-vitro fertilization including 65 with the use of egg donation. Testing options included detection of trisomy 13,18,21 and optional detection of Sex chromosome aneuploidies (SCAs) and fetal sex. The panel offered for SCAs includes 45X, XXX, XXY, XXYY and XYY constitution. Women provided informed consent and maternal blood (20ml) collected in BCT StreckTubes (Streck, Inc, Omaha, NE) and was sent via courier for testing at the CAP accredited, CLIA certified laboratory of NIPD Genetics Public company Ltd. Sample demographics and outcome information was provided by the clinician and was compiled and reviewed to determine the characteristics of this patient population, as well as estimate the assay performance in our clinical setting.
Pre-test and post-test counselling
Patients were provided with a careful and detailed counselling regarding the benefits, risks and limitations of NIPT testing and provided the relevant consent form. NIPT analysis was conducted immediately for each sample and results were delivered using an electronic web-system within 7 days from sample receipt at NIPD Genetics (Nicosia, Cyprus). Women with a positive NIPT were provided the option of invasive testing. Post-test counselling was given to all participants on the basis of their test results.
Invasive testing
For invasive testing chorionic villus sampling (CVS) or amniocentesis was performed. Comparative genomic hybridization (aCGH) analysis was performed using a customized 60K CGX Chip v2 (Perkin Elmer by Agilent Technologies, Inc, Finland) and the data were analysed by a Genoglyphix aCGH software (PerkinElmer, Finland). Banding cytogenetic was performed using routine techniques on G-banded metaphase chromosomes of CVS or cultured amniotic fluid cells. Centromeric probes were used for FISH analysis. Karyotypes were interpreted according to ISCN.
Results
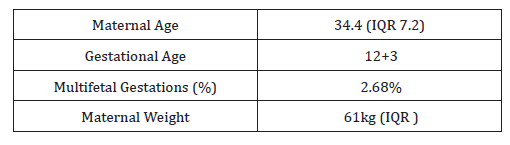
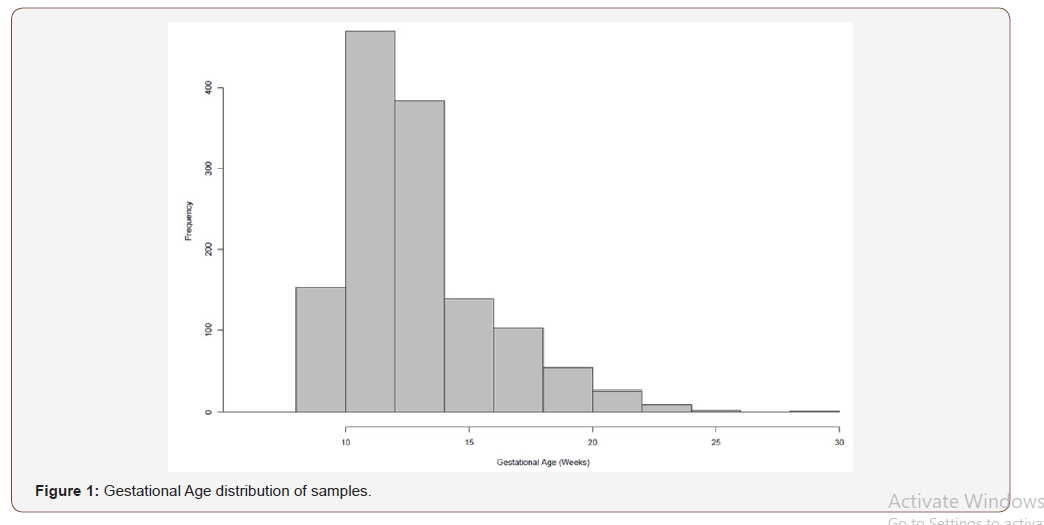
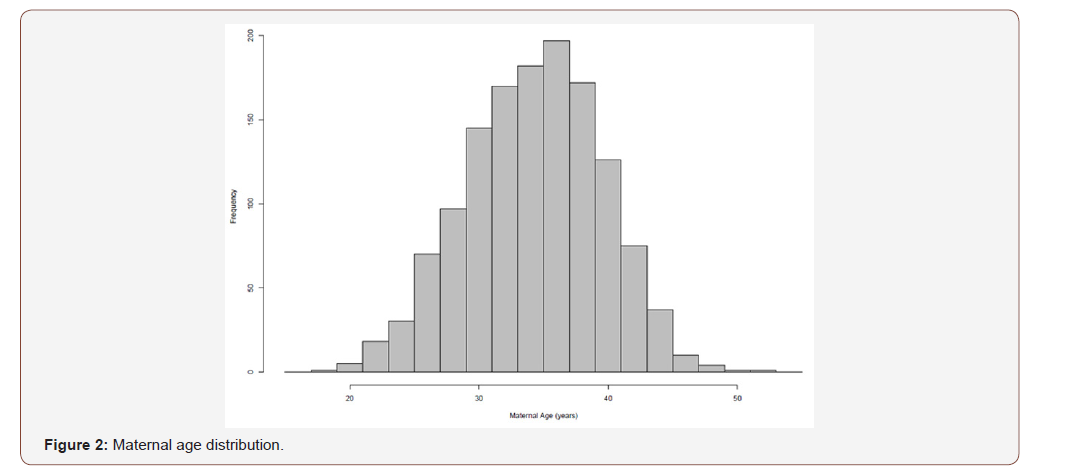
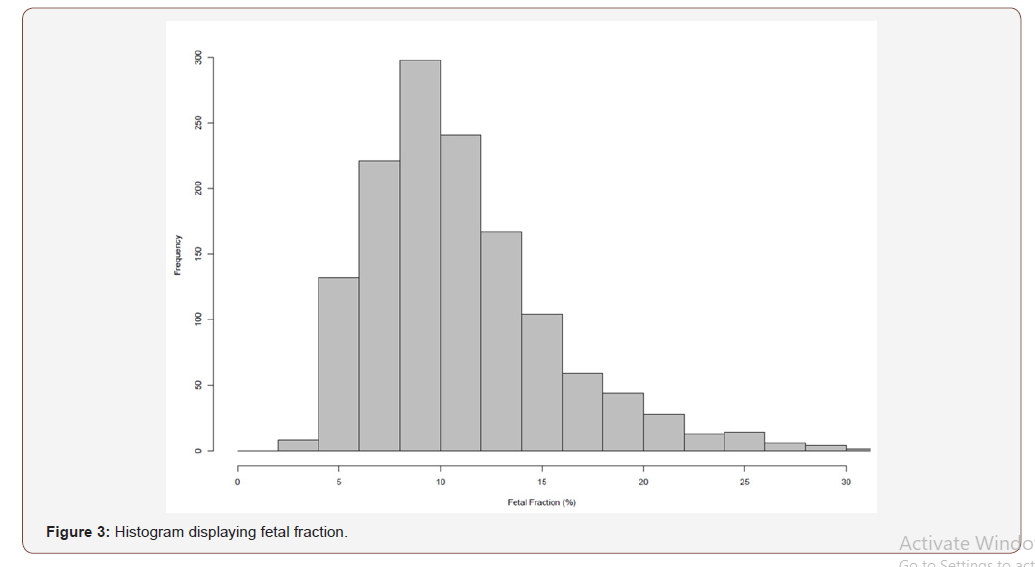
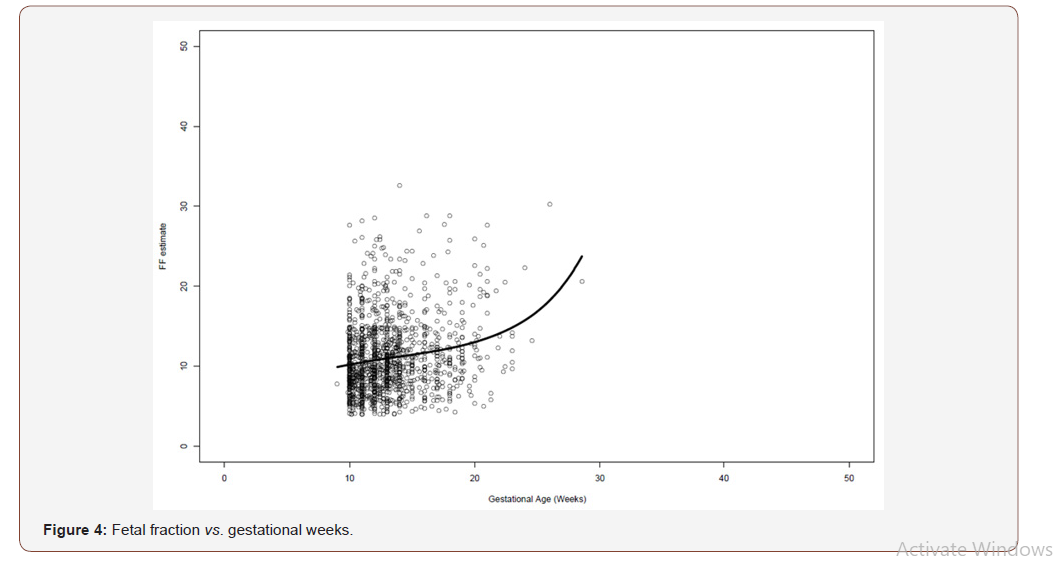
The median gestational age of this patient cohort (n=1382) was 12+3 weeks (Table 1), and the median maternal age was 34.4 (IQR 7.2) years. The median weight was 61 kg (IQR) (Table 1). The overall distributions of gestational weeks and maternal age are depicted in Figure 1 and Figure 2 respectively. In this cohort twin pregnancy samples represented 2.68% of all referrals (Table 1). In this cohort of 1382 cases, 246 cases requested detection of trisomy’s 13,18 and 21, 47 cases requested detection of trisomy’s 13,18,21 and fetal sex and 1089 cases requested the detection of trisomy’s 13,18,21, fetal sex and SCAs. The detection of SCAs was not an available option in twin pregnancies. In this cohort, 1325 cases were singleton pregnancies, 37 were twin pregnancies and 20 were vanished twin pregnancies. The cohort included 169 pregnancies conceived by in-vitro fertilization out of which 65 were performed with the use of egg donation. In the cohort of twin pregnancies, 23 were dichorionic and 14 were monochorionic. The median fetal fraction of reported samples was 10.2% (Figure 3). The fetal fraction increased as gestational weeks increased and exhibited a weak positive correlation (r= 0.19) (Figure 4).
Table 1: Patient Demographics.
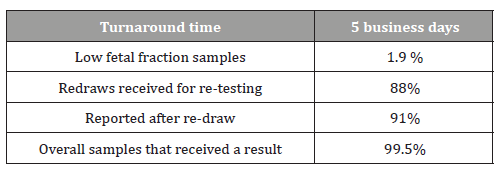
In this sample cohort 99.5% of samples received a result (1375/1382). 1.9% of samples exhibited insufficient fetal fraction and a redraw was requested (27/1382). Specifically, 27 samples exhibited insufficient fetal fraction of which 22 samples were redrawn and sent to NIPD Genetics laboratories for retesting. Twenty out of 22 samples received a result following re-testing (91%). Overall, the median TAT time for reporting was 5 business days (Table 2).
Table 2: Test Performance.
The incidence of trisomy’s 13,18,21 was 1.6% and the incidence of SCAs was 0.5% (Table 3). In summary, 14 trisomy 21 cases were detected by Veracity NIPT. Follow-up information was available for 13/14 cases and all were confirmed (13/13). Four samples were reported as trisomy 18. Follow-up information was available for all samples and all were confirmed (4/4). One sample was reported as trisomy 13 and was confirmed with follow-up confirmatory testing (1/1). Two cases were reported as 45X with no available confirmatory testing information. One pregnancy loss was reported. One sample was reported as trisomy X and was confirmed with follow-up confirmatory testing (1/1). Two samples were reported as XXY and were confirmed with follow-up confirmatory testing (2/2). One case was reported as XYY with no available follow-up information.
Table 3: Summary of NIPT results and follow-up information.
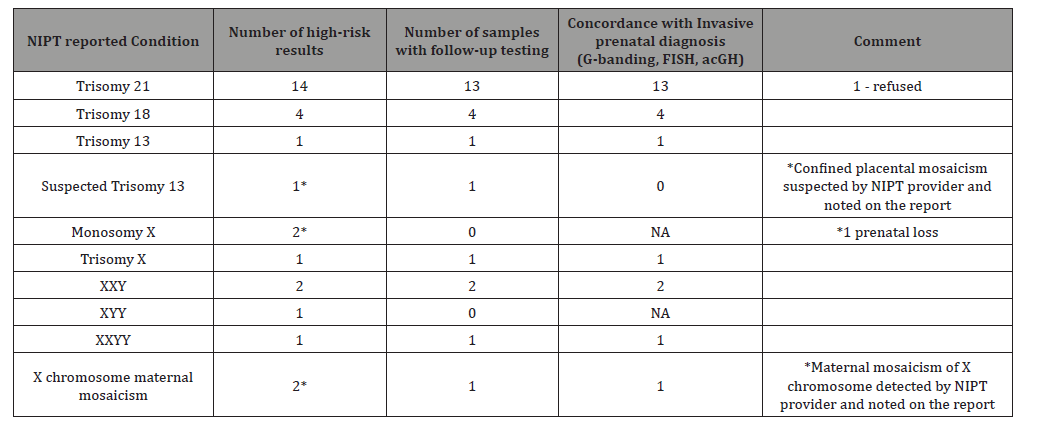
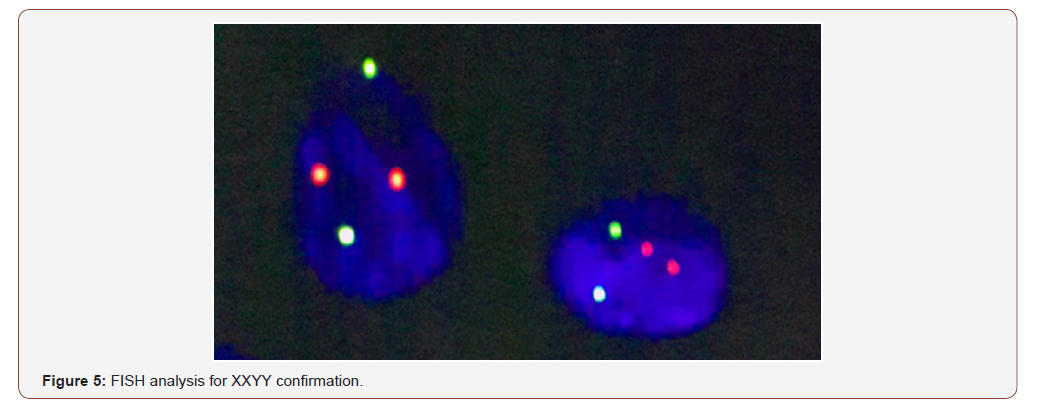
One case was reported by Veracity as suspected trisomy 13 due to boarder line risk scores suggesting the presence of confined placental mosaicism and follow-up testing indicated a normal karyotype result. One case that previously received an inconclusive report using a different NIPT test, was tested by Veracity and was reported positive for XXYY. The Veracity result was confirmed at our centre using FISH analysis (Figure 5) two cases received a Veracity report suggestive of maternal mosaicism of Chromosome X. Follow-up karyotype testing was performed at our centre for one of the two cases using maternal peripheral blood and buccal swab, which confirmed maternal mosaicism.
Discussion
In this study, we evaluated the performance of Veracity under routine NIPT conditions of samples collected in Mother and Child Clinic in a cohort of 1382 mixed risk pregnancies with median 12+3 gestational weeks and 34.4 years of age. The cohort was diverse and included singleton and twin pregnancies, as well as pregnancies achieved with in-vitro fertilization (IVF), and vanished twin and oocyte donation cases and demonstrated high sensitivity (100%) and specificity (100%). Veracity exhibited excellent performance, concordant to previously published validation data and a large scale clinical performance assessment [1,7]. Although tested in a relatively small cohort (n-1382), Veracity NIPT test exhibited broad application prospects for prenatal screening of fetal chromosomal aneuploidies.
Results data were available for 99.5% of samples with TAT of approximately 5 business days. Approximately 2.0% of samples in this study presented a fetal fraction below 4%. We observed that a high proportion of such samples were obtained from participants with relatively higher body weight. This finding is in accordance with previously published data [8,9]. Repeat cfDNA testing was effective in providing reliable results after initial failure due to low ff. In this context it is important that Veracity estimates ff accurately and provides informed recommendations regarding re-testing. Based on this information, our centre utilizes practice guidelines to consider fetal fraction, maternal weight, gestational age and other clinical pregnancy indications in counselling and recommending repeat testing.
In our population, NIPT provided clear actionable results for 99.5% of reported samples. 1 sample received a custom report for suspected trisomy 13 due to boarder line risk score inconsistent with the fetal fraction of 11.5%. This sample was further investigated using amniotic fluid karyotyping and was found to be normal. These results are most likely due to confined placental mosaicism (CPM). In fact, most false positive results for trisomy 13 and monosomy X are a consequence of confined placental mosaicism (CPM). As previously reported, CPM has a higher incidence in monosomy X and trisomy 13 [10-12], which likely explains the lower observed PPV for trisomy 13 and monosomy X in NIPT testing. Two samples received a customized report with an indication of maternal chromosome X mosaicism. Follow-up testing and maternal karyotyping was performed in 1 of the 2 cases and confirmed the maternal mosaicism (45,X/47,XXX/46 XX) in buccal epithelial cells (89/6/5) and peripheral blood (73/20/7). Other reasons for NIPT discrepancies can include an undetected vanishing twin, maternal chromosome abnormality (mosaic or otherwise) and maternal metastatic disease.
Based on our experience the majority of patients in our cohort accepted confirmatory prenatal diagnosis following the international guidelines. We strongly believe that comprehensive pre-test and post-test counselling is very important and should address the importance of confirmatory testing and the benefits of early diagnosis. As such, NIPT can be integrated into current practice in conjunction with appropriate counselling.
In this report we convey a complete and robust clinical picture of the clinical performance of Veracity under routine NIPT testing in our clinic and provide practical information and performance metrics in the Russian population. We will continue to analyse clinical data and explore the use of NIPT and evaluate practicalities and challenges of NIPT implementation in the clinic. Conclusively, based on our experience Veracity is a powerful NIPT test that provides clinicians and prospective parents with important information to make informed decisions regarding pregnancy management.
Acknowledgement
None.
Conflict of Interest
Authors declare no conflict of interest.
References
- Koumbaris G, Kypri E, Tsangaras K, Achilleos A, Mina P, et al. (2016) Cell-Free DNA analysis of targeted genomic regions in maternal plasma for non-invasive prenatal testing of trisomy 21, trisomy 18, trisomy 13, and fetal sex. Clin Chem 62(6): 848-855.
- Chiu RW, Allen Chan KC, Gao Y, Lau VY, Zheng W, et al. (2008) Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci USA 105(51): 20458-20463.
- Fan HC, Blumenfeld YJ, Chitkara U, Hudgins L, Quake SR (2008) Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc Natl Acad Sci USA 105(42): 16266-16271.
- Stokowski R, Wang E, White K, Batey A, Jacobsson B, et al. (2015) Clinical performance of non-invasive prenatal testing (NIPT) using targeted cell-free DNA analysis in maternal plasma with microarrays or next generation sequencing (NGS) is consistent across multiple controlled clinical studies. Prenat Diagn 35(12): 1243-1246.
- (2015) Committee Opinion No. 640: Cell-Free DNA Screening For Fetal Aneuploidy. Obstet Gynecol 126(3): e31-e37.
- Gregg AR, Skotko BG, Benkendorf JL, Monaghan KG, Bajaj K, et al. (2016) Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med 18(10): 1056-1065.
- Kypri E, Loannides M, Touvana E, Neophytou I, Mina P, et al. (2019) Non-invasive prenatal testing of fetal chromosomal aneuploidies: Validation and clinical performance of the veracity test. Mol Cytogenet 12: 34.
- Wang E, Batey A, Struble C, Musci T, Song K, et al. (2013) Gestational age and maternal weight effects on fetal cell-free DNA in maternal plasma. Prenat Diagn 33(7): 662-666.
- Ashoor G, Syngelaki A, Wagner M, Birdir C, Nicolaides KH (2012) Chromosome-selective sequencing of maternal plasma cell-free DNA for first-trimester detection of trisomy 21 and trisomy 18. Am J Obstet Gynecol 206(4): 322.
- Grati FR, Bajaj K, Zanatta V, Malvestiti F, Malvestiti B, et al. (2017) Implications of fetoplacental mosaicism on cell-free DNA testing for sex chromosome aneuploidies. Prenat Diagn 37(10): 1017-1027.
- Grati FR, Malvestiti F, Ferreira JC, Bajaj K, Gaetani E, et al. (2014) Fetoplacental mosaicism: potential implications for false-positive and false-negative noninvasive prenatal screening results. Genet Med 16(8): 620-624.
- Grati FR (2014) Chromosomal Mosaicism in Human Feto-Placental Development: Implications for Prenatal Diagnosis. J Clin Med 3(3): 809-837.
-
Valentina G, Ekaterina K, Yulia T, Mark K. Validation and Clinical Use of the Non-Invasive Prenatal Test “Veracity”. W J Gynecol Women’s Health. 3(5): 2020. WJGWH.MS.ID.000573.
Prenatal Test, Veracity, Fetal aneuploidies, Gestational weeks, trisomy, Sex chromosome aneuploidies, Karyotypes, in-vitro fertilization, Twin pregnancies, Amniotic fluid
-

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.






