 Research Article
Research Article
Multidimensional Classification of Radioprotective Activity of Sulfur-Containing Chemical Agents
Mukhomorov VK*
Physical Department, State Polytechnical University, Saint-Petersburg, Russia
Vladimir Mukhomorov, Physical Department, State Polytechnical University, Saint-Petersburg, Russia
Received Date: September 25, 2024; Published Date: October 08, 2024
Abstract
To establish the relationships between the molecular structure of sulfur-containing agents and their radioprotective effectiveness, methods of factor, correlation and cluster analysis were used. The dose of agents, electronic and informational features of molecules are used as explanatory features. Classification rules were established that separate effective radioprotectors from ineffective agents. To determine the molecular features, it is sufficient to know only the molecular structure of the agent. A structural shift was found for the relationship between electronic and informational molecular features, which differs significantly for effective and ineffective drugs. Factors limiting the radioprotective efficiency of chemical agents are discussed.
Keywords:Modeling; cluster analysis; molecular features; pseudopotential; electronic feature; information function; structural shift; dose; classification rule
Abbreviations:RPA: Radioprotective activity
Introduction
Modeling (or the formation of a classification rule) begins with the identification of objectively existing cause-and-effect relationships, as well as with the establishment of essential features, properties, relationships, and patterns of development of the modeled phenomena, which must be abstracted from secondary, insignificant random factors and relationships. The goal of modeling is defined as the ability to reproduce the characteristics of a certain object on other objects specially created to study these characteristics. The main requirement for a model is that the model should be some likeness of the phenomenon (object) being studied. However, it should be borne in mind that the statistical modeling apparatus alone does not make it possible to make an unambiguous judgment about the presence of cause-and-effect relationships in models. The validity of cause-and-effect relationships is determined by the correctness of the task, the correct choice of the model type, the methods of selecting factor features and, first of all, the qualitative, theoretical analysis of the modeling object. Preliminary qualitative analysis is the starting point for statistical modeling.
The goal of modeling the relationship between the structure of molecules and their bioactivity is not to replace the knowledge, ideas and experience of biologists, physicians and chemists, but to obtain a tool that will allow predicting the properties of new, as yet unstudied chemical compounds. On the other hand, modeling will allow us to identify or suggest possible biochemical processes in which an exogenous substance may participate. One of the pressing problems of modern chemistry of biologically active substances remains the problem of creating drugs that are effective in terms of radioprotection. The main requirements for effective drugs are small doses, low toxicity, and absence of side effects. An important aspect of modeling is the accessible acquisition of molecular features that are associated with the biological action of drugs. Since the effect of interaction of drugs with the biosystem depends on many conditions, the model has a multidimensional nature. Therefore, when searching for dependencies, it is preferable to use multidimensional models.
Methodology
To model the biological action of agents, new molecular descriptors are proposed here that characterize the electronic and informational properties of molecules. The quantitative approach proposed here for determining molecular descriptors requires knowledge of only the structural formula of chemical compounds. Thus, the value Z, which is associated with the pseudopotential of the molecule, will be used as an electronic property of the molecule:

Here ni is the number of atoms of the i-th type. The Zi attribute determines the number of electrons in the outer electron shell of the i-th atom. The summation is performed over all atoms in the molecule; N is the total number of atoms. That is, the Z feature takes into account the average number of electrons in the outer shell of atoms in a molecule. It is known [1-3] that the Z feature, which characterizes the number of electrons per atom of electrons, is a factor in the quantitative determination of pseudopotential. The model pseudopotential of the core ions of the molecule is weakened compared to the Coulomb field of an isolated ion due to screening by external (valence) electrons

where f (r) and F(r) are corrections to the Coulomb potential that depend on the distance r between the core of the molecule and the electron; e is the charge of the electron, RM is the radius of the scattering center.
Determining the actual molecular potential involves complex quantum calculations. This significantly complicates the construction of a practically convenient model for obtaining a rule for classifying agents. At the same time, it is known [1-3] that the electronic feature Z allows for the reliable reproduction of many properties of condensed media. For example, a model pseudopotential correctly reproduces the nature of electron scattering on atomic potentials in a solid. The model molecular pseudopotential method takes into account only the electrons in the outer (valence) shell of the scattering center. As is known, the chemical and biological properties of molecules are determined precisely by the electronic state of a relatively small group of outer valence electrons. The remaining electrons of the atoms (the socalled core electrons) have almost no effect on the physicochemical and biological processes in which the molecule participates.
In addition to the electronic feature Z, molecular information features will also be used, which characterize the complexity (diversity) of the molecule structure. First of all, this is the Shannon information function [4], which for a discrete data set is defined as follows:

Sometimes function (3) is called entropy, since the information feature has the same mathematical form that is used in thermodynamics or statistical mechanics. In formula (3) the notation pi=ni/N is used. For the relations pi=ni/N the following conditions are met: 0≤ pi≤ 1, pi =1 , and the condition pi=0 determines the impossibility of the i-th event. The ratio ni/N determines the fractional participation of the i-th type of atom in the molecule. However, in fact, instead of the probabilities that were used by Shannon, in formula (3), Kolmogorov’s combinatorial representation [5] is used about the share participation of elements of a set in a common set (the elements of a set are the atoms in a molecule).
The information function quantitatively characterizes the measure of chaos, that is, the measure of disorder of a multicomponent system. The larger the value of H, the greater the uncertainty of events. The information function is zero if the probability of any event is 1 and the probabilities of all other events are zero. The information function H is an integral characteristic of a molecule that gives a measure of the uncertainty (or diversity, or ignorance) of the molecular structure of a chemical compound. Function (3) takes on a maximum value when the values of pi are equal for all events. That is, the object is homogeneous. A uniform distribution corresponds to the maximum uncertainty of the object’s state. The smaller the value of the function H, the more diverse the multicomponent system. In addition to the full information function H, the partial information function dH will also be used:

Here pC,H =NC,H / N , NH is the number of hydrogen atoms, NC is the number of carbon atoms.
Table 1 shows two independent samples of effective radioprotectors and inactive sulfur-containing chemical compounds. The first group (marked “+”) contains agents that have effective radioprotective activity (RPA) (dose ≤ 1 mM/kg; survival rate greater than 50%). The second group (marked “–“) includes agents that do not have effective antiradiation activity even in relatively large doses used (dose > 2 mM/kg). Table 1 shows the values of the electronic feature Z for each molecule, as well as the informational molecular descriptors H and dH. Using a sample of 100 chemical compounds that are random and independent, we will determine the average values of the Z feature for effective radioprotectors Z1av (number of chemical compounds N1 = 57) and ineffective Z2av (number of chemical compounds N2 = 43) chemical compounds. Knowing the average values of independent random variables allows us to identify relationships that are characteristic of mass phenomena. The following unbiased and consistent estimate [6] of the sample or empirical simple mean for a discrete variable Z was obtained:
N =100, Zav = 2.83± 0.03CU is the sample mean; reliability of the mean value: t =94>t0.05cr (f=N − 1)= 1.982; (2.77−2.88) there is a 95% two-sided confidence interval; reliability of the mean value

is an empirical standard; Kolmogorov-Smirnov normality test:

David-Hartley-Pearson test:

For the set of effective radioprotectors, the following statistics of the electronic signature of the 1 Z molecule were also obtained:
N1 = 57, Ziav=2.689±0.03 CU, reliability of the mean value: t =81 > t0.05cr (f = N1 −1) = 2.004 ; 95% two-sided confidence interval: (2.62 − 2.77) ; Z1 min=2.20, Z1max= 3.300 ,
empirical standard SZ1=0.251 Kolmogorov-Smirnov normality test: dmax=0.07, λ=0.53< λ 0.8cr=1.07 ,p=0.37>α= 0.05; David- Hartley-Pearson test:

It follows that the true value of the arithmetic mean is with a probability of 0.95 in the range of values 2.66 < Z1av< 2.72 conventional units. For the second group of agents (ineffective or weakly active), the following statistics were obtained:
N2 =4.3, Z2 =2.99 ±0.04, reliability of the mean value: t =70>t0.05cr (f =N2− 1)= 1.994; 95% two-sided confidence interval: 2.90 − 3.08 ; Z2min = 2.316 , Z2max = 3.818 , empirical standard: S2Z= 0.284 , Wilk-Shapiro homogeneity test: W =0.977> W0.05cr (N2)= 0.923 ; Kolmogorov-Smirnov normality test: d max = 0.0963, λ = 0.63< λ 0.08cr = 1.07 , p = 0.55 >α = 0.05 ; David-Hartley-Pearson test:

According to the confidence interval, the true mean value of the feature Z2 is within the range of: with a probability of 0.95 : 2.94< Z2av< 3.04 . Therefore, the interval estimates for Z1av and Z2av do not overlap.
A set of elements can be considered independent only if its average value is statistically significantly different from the average values of other groups. That is, it is necessary to check whether these average values really belong to different sets of chemical compounds. Let us accept as the null hypothesis the equality of the mean values of two random samples Z1 and Z2 . The alternative hypothesis is the following: Z1av ≠ Z2av. Determining the significance of the discrepancy between average values is important because it allows us to establish whether their differences are random or not. First, we will determine the difference between the variances of SZ12 and SZ 22 using the F-ratio. To do this, we compare the ratio of the larger variance to the smaller variance at a significance level of α = 0.05 and degrees of freedom f1 and f2 [7,8]:

That is, the empirical F-value is less than the tabulated value with 95% reliability. Therefore, the difference in dispersions can be considered insignificant. Therefore, to assess the significance of the difference between two sample mean values of the features Z1av (6) and Z2av (7), we can use the student-fisher test [9]:

Here S2 = 0.0859 is the variance for the total population (N =100) ; the tabular value of the student’s t-distribution t0.05cr (f ) has f = N1 + N2 − 2 degrees of freedom and a significance level of α = 0.05; N = N1 + N2. From inequality (9) it follows that the null hypothesis is rejected at a significance level of α = 0.05. The difference in mean values cannot be explained by random deviations with a reliability of 0.95. Since inequality (9) is satisfied, there is a relationship between the dependent feature RPA agent effectiveness and the categorical feature Z. Effective radioprotectors have a Z-sign that is, on average, greater than the Z-sign of ineffective drugs. That is, it is necessary to accept the hypothesis that RPA effective drugs are most likely grouped around the average value of Z1av, and ineffective drugs are grouped around the value of Z2av.
Additional information about the significant difference between Z2av and Z1av can be obtained by determining the biserial correlation coefficient. The biserial correlation coefficient is useful when one of the two variables (in this case RPA) is dichotomous (takes qualitative values “+” or “–”), and the second variable Z is measured quantitatively (strong scale). The point-biserial correlation coefficient is calculated as follows [10]:

The following results were used here:
N1 =57, N2 =43, N =100, Z1av =2.689, Z2av=2.989 Sz =0.293 is the standard deviation for the original sample. The significance of the point-biserial correlation coefficient is tested using the following inequalities [11]:

Since the empirical values of t and F are greater than the tabular values (these inequalities are also observed at significance α = 0.999) , the null hypothesis that the correlation coefficient is equal to zero can be rejected. Therefore, there is a significant difference between the average values of the electronic feature Z for the RPA of effective agents and inactive (or weakly active) agents. The value of the electronic feature Z ≤ Z av = (2.83± 0.03) CU (sample size N = 100) (5) for most chemical compounds that were used in a low dose (dose D ≤ 1 mM/kg). This value of the electronic feature can be considered as a threshold or boundary value: Zthr ≡ Z av . At the same time, for chemical compounds that do not have an RPA effect (even when used in very high doses D > 2 mM/ kg), the characteristic is usually Z > Z av . The classification rule assumes that the antiradiation effect of the agent does not occur at any value of the Z indicator, but mainly when the indicator becomes less than a certain threshold value.
Since only one molecular feature was used, it can be assumed that an objective pattern was identified. Next, a formal comparison of the frequencies of occurrence of various atoms in the effective chemical compounds and ineffective preparations of the RPA was performed (Table 1). Figure 1A shows the frequency of occurrence of carbon, hydrogen, nitrogen, oxygen and sulfur atoms in effective radioprotectors. Ineffective sulfur-containing chemical compounds are shown in (Figure 1B). Such graphic information allows one to obtain some approximate idea of the hypothetical gross formula of the relay protection and automation equipment of an effective radioprotector. Using the data in Table 1, we can indicate the most frequently encountered atoms (in the gross formula of the molecule) contained in a hypothetically effective radioprotector and satisfying the principle of homogeneity (Figure 1A): phosphorus (P) ~ 1 atom, sulfur (S) ~ 1 atom, nitrogen (N) ~ 2 atoms, oxygen (O) ~ 3 atoms, carbon (C) ~ 5 – 6 atoms, hydrogen (H) ~ 17 atoms.
Table 1: Radioprotective activity (RPA) [6-12] of sulfur-containing chemical compounds, the dose of the drug used, as well as the electronic and information factor-signs of molecules.

*) The number of electrons in the outer shell of an atom: Z(H) = 1, Z(C) = 4, Z(N) = 5, Z(S) = 6, Z(P) = 5, Z(O) = 6, Z(Pb) = 4, Z (Br, F) = 7.
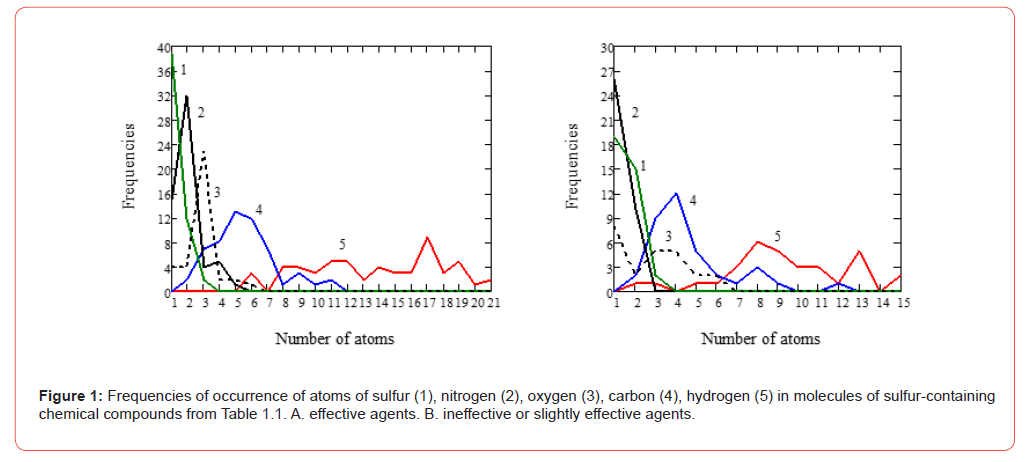
It can be noted that this sequence of the number of atoms is close to the sequence of the first numbers of the Fibonacci series: 1, 1, 2, 3, 5, 8, 13. The peculiarity of the sequence of these numbers is that each of its members, starting with the third, is equal to the sum of the two previous ones. The Fibonacci series is widely found in science, technology, art and nature. Assuming these frequencies to be optimal, we obtain the value of the electron factor Zeff = 2.62 − 2.67 CU, which is close to Z1av =2.689 CU (6). For ineffective or weakly effective chemical compounds, the most probable distribution of atoms in a hypothetical molecule is as follows: P ~ 1, S ~ 2, N ~ 1, O ~ 1, C ~ 4, H ~ 8 - 10 Zneff = 2.84 – 3.06 CU ≈ Z2 av = 2.989 CU (7). Comparing Figures 1A and 1B, one can notice that the frequencies of occurrence of the hydrogen atom often have different directions in effective radioprotectors and ineffective chemical compounds.
From the point of view of possible biochemical processes, it is possible that low values of the molecular characteristic Z contribute to the capture of hydrated electrons (eaq) by the radioprotector molecule, which are a product of water radiolysis. Hydrated electron is a strong reducing agent. It is known that the effect of radiation on water leads to the formation of highly active radicals OH, H, H2 O2 and eaq. The hydrated electron is capable of reacting with many dissolved substances present in quantities of several hundred micromoles [12]. The possibility of the appearance of a very strong radical (eaq)2 as a result of intense irradiation cannot be ruled out [13]. A similar test of significant difference in mean values can be used for the information features H and dH. For RPA of effective agents, the average feature values are as follows: H1 av = 1.749 bits, dHav = 0.0021 bits. For chemical compounds that do not have radioprotection, the average values of the features are respectively: H2 av = 1.866 bits, dH2 av = -0.0087 bits. Thus, RPA effective chemical compounds and ineffective drugs differ in all three explanatory molecular features.
Results and Discussion However, it is important to integrate many different, simultaneously acting explanatory factors into one model. Multivariate cluster analysis provides such an opportunity. The advantages of cluster analysis over other statistical methods are that it does not require the preliminary creation of a training model, the set of initial data can be arbitrary and is not necessarily normally distributed. In multivariate cluster analysis [14], classification is carried out according to four features that simultaneously characterize agents (dose, Z, H and dH). The main task of cluster analysis is to divide all objects into a certain number of optimal subsets within a common set of initial data. Cluster analysis is a reliable statistical tool for multivariate data analysis that allows one to identify not always obvious relationships and groupings in large volumes of information. The purpose of multidimensional classification is to identify rules (or classifications) by which the analyzed objects are divided into several structurally different groups.
Thus, cluster analysis forms homogeneous groupings, but which are heterogeneous from each other. The assignment of an object to a particular group is defined as its identification. Multivariate cluster analysis can help uncover hidden patterns and structures in data that are not obvious at first glance. Cluster analysis considers n objects that are described by m heterogeneous features. Each object is represented as a point in m-dimensional space. As a result of the cluster analysis, all objects are divided into homogeneous groups, with each object belonging to one, and only one, subset of the partition (cluster). To determine the distance between subsets of objects, the concept of metric d is introduced. For quantitative features, the Euclidean distance between clusters in a multidimensional space is determined from the relation

Here Xik is the value of the k-th feature on the i-th object. Standardization (norming) of variables is usually performed beforehand for populations that differ significantly in size.
This allows you to bring all values of the transformed features to a single range. However, it should be borne in mind that with a significant difference in the dimensionality of variables, the standardization operation reduces the accuracy of clustering. It should also be noted that if cluster analysis is preceded by statistical analysis of features, then the sample does not need to be adjusted. The selected features should be informative (that is, have essential properties for classification) and should not strongly correlate with each other. The test showed that the features Z and H, as well as H and dH, are interrelated. The correlation coefficients are 0.87 and -0.73, respectively. For the cluster analysis method, the relationship between features is considered significant if the correlation coefficient |R| > 0.7 [15]. Therefore, one of the features, in this case H, can be excluded from the analysis. This reduces the dimensionality of the multidimensional space. In addition, the sets of each feature must be homogeneous, since the cluster analysis method is very sensitive to outliers.
Multivariate cluster analysis differs from traditional statistical methods in that the classification method does not require any restrictions related to statistical criteria. Application of the multivariate cluster analysis method to the data in Table 1 allowed us to identify four clusters. Within each cluster, objects are maximally similar to each other, while between clusters there is minimal similarity. In general, if an object characterized by m features is represented by a point in n-dimensional space, then the similarity of objects to each other will be determined by the value 1/(1 + d). The larger this value, the closer the compared clusters are to each other. Euclidean distances (Table 2) were obtained for objects from Table 1. Euclidean distances allow us to divide populations into homogeneous groups based on four characteristics simultaneously. The distance between clusters shows how different objects are in different clusters. Obviously, the larger the metric d, the more significant the difference between objects.
The k-means algorithm proposed by G. Steinhaus [16] allows for the division of objects into clusters in such a way that the objects within the clusters are as close to each other as possible (homogeneous). At the same time, objects from different clusters are maximally different (heterogeneous). Moreover, each object belongs to one and only one cluster. Cluster analysis divided all chemical compounds into four different groups (Table 2), which included the following numbers of objects (agents): 38, 22, 27 and 13. Cluster analysis of the initial data showed that the average values of the Z feature for RPA effective compounds (clusters 1 and 2) are below the range (2.706 – 2.715) conventional units. At the same time, the information feature dH for effective agents should not be lower than the range of values (-0.002 ÷ 0.0074) bits. The obtained range of the electronic feature is below the expected threshold value Zthr = 2.83–2.87 CU. For ineffective agents, cluster analysis yields the following average values of the Z feature: 2.97 conventional units (cluster 3) and 2.99 conventional units (cluster 4).
These values a re greater than the expected threshold value Zthr and are close to the value Z2av = 2.99 CU (7). For ineffective drugs, the information factor dH remains negative and relatively large in absolute value. The k-means algorithm remains one of the main tools in cluster analysis due to its relative simplicity and the speed of the results obtained. Thus, cluster analysis showed that RPA effective drugs in a low dose (for example, cluster 2) should have a relatively high average value of the Z feature (≈ 2.8 conventional units) and a dH feature value > -0.0020 bits. In addition, a set (cluster 1) of effective drugs with a relatively low electronic signature Z ≈ 2.72 bits is possible. However, for such agents, the dH feature should be positive and have a relatively large value of ≈ 0.0074 bits. This cluster is formed near the “dose” cutoff of ≈ 0.9436 mM/kg, which separates RPA effective compounds from ineffective agents. That is, a decrease in the value of the electronic feature Z of the molecule is accompanied by an increase in the value of the information feature dH to its positive value.
At the same time, an increase in the Z feature (for example, for cluster 4 preparations to a value of 2.97 CU) is accompanied by a significant decrease in the information feature dH to a value of ≈ -0.0107 bits. This reduction is associated with the absence of a radioprotective effect. Multivariate analysis provides a more detailed result than using only threshold or mean values of Z and dH features. Cluster analysis identified two potential regions (clusters 1 and 2) for effective radioprotectors. Let us check the interrelationship between the features dH and Z of the two extreme clusters, namely, 1 and 4. For RPA effective agents (cluster 1) there is a significant linear relationship between these features:

sum of squares of residuals: Σ = 0.0212 ; test of adequacy of the linear model [17]:

As calculations have shown, for ineffective agents (cluster 4), such a relationship is absent: R = 0.03. Therefore, a structural shift is observed for the relationship between the dH and Z features when moving from effective to ineffective agents. That is, the drugs in clusters 1 and 4 are separated from each other not only by the values of the dH and Z features, but also by the difference in the internal structure of the molecules, which is expressed in the significance of the relationship between these features. In the cluster analysis methodology, there are no criteria that determine the quality of classification or the number of clusters required for classification. The point is that during classification there may be losses of objects that are not included in any cluster. It is known that the more unidentified objects, the lower the classification quality. From Table 2 it follows that there are no unidentified objects. That is, the total set of objects is equal to the sum of objects in clusters. Consequently, no atypical objects (outliers) that cannot be assigned to any cluster were detected.
Thus, cluster analysis revealed important organization among a variety of chemical compounds. Four homogeneous clusters were found that differ from each other. It is recommended to investigate the idea of the quality of the classification by comparing the results of the original sample with the results of a random sample. Let’s check the representativeness of the analyzed initial sample. The results obtained can be verified by drawing up a random sample. The purpose of analyzing a random sample is to determine whether the sample is representative of the original population. Using a table of random numbers [18], we will compile a partial sample from the original data (Table 1). The random number method allows one to avoid subjective approaches, bias or prejudice in the selection of sample elements, and also to avoid systematic and unintentional errors in the selection of the sample. This random selection serves as an additional test of the null hypothesis. As a result, the following sequence of random numbers was obtained (N10 = 40):

The sequence of random numbers (13) was obtained by selecting numbers (only numbers ≤ 100 were taken into account) starting from the first number of the reference table of random numbers [19]. Thus, all random numbers not exceeding 100 were written out. After matching these random numbers with the drug numbers from Table 1, a random sample of 40 elements was obtained (13). The number of objects in the sample is random, has the same probability and is unbiased. In such a random sample, the sequence of chemical compounds is objectively represented. The statistics for a random sample will be as follows:

The representativeness of the sample for assessing the reliability of the arithmetic mean Z1 av can also be assessed using the following relationship [20,21]:

or using the formula

The following notations are used here: y1= Σi Zi – the sum of the variant series, y2 = Σi Zi2 – the sum of the squares of the variant series. Since the value of Θ is greater than three and the value of Ψ is greater than nine, the sample size is sufficient for the reliability of the arithmetic mean of the population. The significance estimate Θ is also determined using the tabular value of the Fisher distribution:F0.05cr (f1=1;f2 = N10 −1) =4.08 . Let us check the statistical significance of the difference in the mean values Z av = 2.85 (5) and Z10av =2.79 (14) for sample sizes N = 100 and N10 = 40, respectively. To do this, we use relations (8) and (9). The inequality for the ratio of variances F =1.30 < F0.05cr(f2=98;f1=38)=1.60 allows us to use test (9): ΔZ= |Zav−Z10av|=0.04< tav=0.104. Therefore, there is no statistically significant difference in the mean values of two samples that have significantly different sizes.
Now we will present statistics for effective drugs and inactive chemical compounds of a random sample. Statistics of the set of chemical compounds for which the dose D ≤ 1 mM/kg:


a set of chemical compounds for which the dose D > 2 mM/kg: N21=16, Z21av= 2.89±0.06, reliability of the mean value: t =48 > t0.05cr (f=N21−1)=2.120 ; 95% confidence interval (2.75 − 3.02) ; Z21min= 2.316, Z21max = 2.263 , empirical standard:

Additional information about the adequacy of the model can be obtained using the residual sample after excluding random sample elements from the total sample (13): N30 = 60, Z3 av= 2.82 ± 0.04, S3 = 0.257; N31 = 33, Z31 av = 2.63 ± 0.04, S31 = 0.241 (effective agents); N32 = 27, Z32 av = 3.05 ± 0.06, S32 = 0.290 (inactive agents). For this residual sample, as well as for the random sample (13) and the original sample (5), the sequence of mean values is preserved: Z31 av < Z3 av < Z32 av. Using relations (8) and (9), as well as the test [22], it can be shown that there is no statistically significant (at a significance level of α = 0.05) difference in the mean values for samples of different sizes. For example, for samples N10 = 40 (Z10 av = 2.79, SZ10 = 0.257) and N30 = 60 (Z3 av = 2.82, S3 = 0.257), the estimate of the difference in mean values is insignificant:

Test (17) – (18) is applied to samples with non-significantly different or equal variances. Thus, statistical results obtained for significantly different sample sizes can be considered stable. These results do not contradict statistics (5) and (6), and the studied feature Z is represented in the same proportion as in the original sample (N = 100). The groups are homogeneous and, according to the Wilk-Shapiro and David-Hartley-Pearson tests, have a normal distribution. That is, it can be assumed that there is statistical stability (consistency) of the classification. Any random removal of an element from a population does not lead to a significant change in the characteristics of the population of independent elements. The application of multivariate cluster analysis to a random sample (13) leads to practically the same proportions for the content of objects in clusters as for the original sample (indicated in brackets): clusters 1 - 4 contain 17(38), 7(22), 14(27) and 2(13) objects, respectively.
There is a correlation between the number of objects in clusters for two different samples (correlation coefficient 0.96). The Euclidean distances of two different samples are also comparable. For example, the minimum and maximum distances between clusters are close to each other: dmin = 0.4113 (N = 100) and dmin = 0.3288 (N10 = 40), dmax = 5.724 (N = 100) and dmax = 5.8213 (N10 = 40), respectively. The average values of the electronic feature Z for effective drugs Zav = 2.7117 CU (N = 100), Zav = 2.8036 CU (N = 40) and for ineffective drugs Zav = 2.9631 CU (N = 100), Zav = 2.9774 CU (N = 40) are comparable with each other. Similar relationships hold for the average values of the dHav feature. For effective drugs dHav = -0.002 (N = 100) bits and dHav = -0.009 (N = 40) bits. The mean doses for all four clusters for the original sample (N = 100) and the random sample (N = 40) are almost identical. We also note that for both the original sample (Table 1) and the random sample (13), no unidentified objects were detected.
The structural shift (12) of the dH(Z) relationship is also preserved for a random sample. For effective agents (clusters 1 and 2) there is a significant (R = 0.85 > R0.05 cr (f = N1 + N2 – 2 = 22) = 0.404) correlation relationship dH(Z), whereas for ineffective agents such a relationship is absent. Thus, a significant reduction in the original sample size due to the random retention of a relatively small number of representatives of each class did not lead to a significant change in the results of the cluster analysis. For a random sample, the average distance between clusters is 3.8 CU, which is greater than the average distance of 3.2 CU obtained for the original sample. That is, the quality of clustering is even somewhat higher. Using test (8), it can be shown that the average values Z1 av = 2.69 (6) and Z11 av = 2.73 (16), Z2 av = 2.99 (7) and Z21 av= 2.89 (16), as well as Zav = 2.83 (5) and Z10 av = 2.79 (14) do not differ significantly. Therefore, the classification results for the two samples are consistent. For example, we obtain the following inequalities for the RPA of effective agents Z1av and Z11av :

Inequalities (19) and (20) indicate that there is no statistically significant difference between the mean values. Using relations (10) and (11) one can also check the significance [22] of the biserial correlation coefficient rpb= 0.309:

From inequalities (21) it follows that at the 95% confidence level, the average values of the Z feature are statistically significantly higher for agents with RPA effective radioprotective action than for inactive compounds, and their difference is not accidental. Let us check the classification rules for chemical compounds that were not included in the original sample (Table 1).
Such agents include, for example, 2,6-dimethylphenoxytetramethylenaminoethyl thioseric acid: (CH3)2 C6 H3 O(CH2)4 NH2CH2CH2SSO3H. This agent is an effective radioprotector. (dose: 0.3 mM/kg; Z = 2.69 UC < Zthr; dH = 0.04 bits, H = 1.64 bits, RPA = 100%). However, analogues of this compound do not exhibit radioprotective effect [10]: C6H5-S-(CH2)4NHCH 2CH2SSO3H (Z = 2.84 CU ≈ Zthr, dH = 0.025 bits, H = 1.74 bits) and C6H5-SO2-(CH2)4NH-CH2CH2SSO3H (Z = 2.89 CU > Zthr, dH = -0.004 bits, H = 1.81 bits).
These results are consistent with the formulated classification rules of cluster analysis for the molecular features Z and dH. Electronic Z and information dH features were also determined for a number of other sulfur-containing compounds that exhibit effective RPA when used in low doses: 1) disulfide of betamercaptoethylamine (dose: 0.9 mM/kg; Z = 2.69 CU. < Zthr, dH = 0.002 bits, H = 1.64 bits); 2) (CH3)2N-CC6H5-CH(OH)-S-CH2CH2NH2 (dose: 0.88 mM/kg; Z = 2.50 CU < Zthr, dH = 0.0576 bits, H = 1.54 bits); 3) cyclo-C8 H15 O-(CH2)5-HNCH2CH2SH (dose: 0.15 mM/kg; Z = 2.20 CU. < Zthr, dH = 0.105 bits, H = 1.29 bits); 4) AcNHCH2CH2-S-S- (CH2)4-SO2 Na (dose: 0.58 mM/kg, Z = 2.68 CU < Zthr, dH = 0.0036 bits, H = 1.94 bits); 5) C6 H13 CH(CH3)-NH-CH2CH2SSO3H (dose: 0.28 mM/kg; Z = 2.79 CU < Zthr, dH = 0.0036 bits, H = 1.94 bits); 6) cyclo- C6 H11-CH(CH3)-CH2CH2CH2-NHCH2CH2S- SO3 H (dose: 0.08 mM/kg; Z = 2.48 CU < Zthr, dH = 0.064 bits, H = 1.54 bits).
Apparently, the dose, molecular features Z and dH for these chemical compounds satisfy the acceptable ranges formulated by cluster analysis (Tables 2 and 3). For chemical compounds with a feature value Z > Zthr, for example, 1,2,5-thiadiazole-3,4- dicarboxylicacid (dose > 4.60 mM/kg; Z = 4.36 CU > Zthr, dH = 0.040 bits, H = 1.99 bits); 1,2,5-thiadiazole-3 –carboxylic acid (dose > 7.69 mM/kg; Z = 4.20 CU > Zthr, dH = 0.0567 bits); H2 NSSO3 H (dose: 4.65 mM/kg; Z = 3.90 CU > Zthr, dH = -0.5288 bits, H = 1.8468 bits) no effective RPA was found. Apparently, for this group of agents, the electronic feature Z is a factor that limits their RPA effectiveness.
Table 2: Average values of features and cluster content.

Table 3: Average feature sizes and cluster content.

The method used for classifying drugs allows for a rapid assessment of the expected radioprotective effect of chemical compounds in significantly different doses. However, the obtained results cannot claim to be a complete explanation of the causeand- effect relationships between the radioprotective effect of the drug and its molecular structure. A more detailed study of the molecular structure of the molecules is required. For example, for the optical isomers of cysteine and iso-cysteine, the molecular features Z, H and dH will be the same, whereas in the first case the compound has a radioprotective effect, and iso-cysteine does not have a radioprotective effect. When equal doses of S-β- aminobutylisothiouronium are used, the D-form of this compound is more bioactive in terms of radioprotection than the L-form, although both isomers do not differ in toxicity and have the same values of the molecular features Z, H and dH. It is also important to note that some chemical compounds, when used in low doses and for which the inequalities Z ≤ (2.706 – 2.715) conventional units, dH > (-0.002 – 0.0074) bits (Table 2) are satisfied, nevertheless do not have a radioprotective effect.
This result may be due to various possible mechanisms limiting the potential biological activity of drugs. One of such possible mechanisms is discussed in [23]. It has been shown that only with well-defined hydrophobic properties of chemical compounds do optimal transport properties of molecules appear, facilitating the maximum manifestation of biological activity. Let us consider one of the possible mechanisms for limiting the biological activity of drugs in more detail. Let there be a homologous series of chemical compounds CH3 (CH2)nNHCH2CH2SSO3H, where the index n = 0. It is known [24] that if the index n takes values in the intervals: 0–5 and 13–17, then these drugs do not have effective radioprotective activity. A similar situation occurs for N-substituted S-2-aminoethyl thiosulfates, which have radioprotective activity in the range of index values n ≈ 6 - 12 (Z = (2.37–2.56) CU, H = (1.46–1.63) bits, dH = (-0.069 – 0.0927) bits). The maximum toxic properties (LD50) of these molecules appear in almost the same range of n index values. The dependence of biological activity A (result feature or target function) of chemical compounds of the series CH3(CH2)nNHCH 2CH2SSO3H on their hydrophobic properties can be approximated by the following analytical form [25, 26].
Here Δπ = Σi πi is the sum of additional additive contributions
to the hydrophobicity of the molecule, caused by the addition of
CH2 atomic groups. Hydrophobicity π determines the processes
of agent distribution in the biosystem. The physicochemical role
of distribution consists of the ability of a molecule to penetrate to
the sites of its action in a biosystem. The additional contribution
of substituents to the hydrophobicity of the original molecule is
determined by the method of additive increments. The contribution
of each group of CH2 atoms to the hydrophobicity of the molecule is
taken to be equal to: 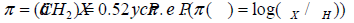 [27]. PH is the distribution coefficient in the octanol-water system
for the original unsubstituted molecule. PХ is the distribution
coefficient for a molecule with substituent X. Nonlinear regression
(22) is characterized by the presence of a maximum. The
nonlinearity is due to the fact that drugs with a high π value are
delayed in their movement in the body due to binding to lipophilic
sites, whereas very hydrophilic molecules can be trapped in water
pockets [28]. In the specialized literature it has been noted that the
biological effect of some substances is significantly reduced with an
increase in their solubility in water. The competition of these two
processes leads to the existence of some optimal (in the region of
the maximum) hydrophobic properties of chemical compounds. In
the range of values n = 9 – 14 (additional hydrophobicity of the base
molecule is equal to Δπ ≈ 5 – 7 conventional units), the toxicity of the
drugs reaches its maximum value. Their maximum radioprotective
effect is also manifested in this same range of n index values. Thus,
although chemical compounds with index n > 14 have values of Z
< (2.706 – 2.715) CU, as well as dH < (-0.069 – 0.0922) bits and
are probably potentially active radioprotectors, nevertheless, their
radioprotective effect is limited by the hydrophobic properties of
the molecules.
[27]. PH is the distribution coefficient in the octanol-water system
for the original unsubstituted molecule. PХ is the distribution
coefficient for a molecule with substituent X. Nonlinear regression
(22) is characterized by the presence of a maximum. The
nonlinearity is due to the fact that drugs with a high π value are
delayed in their movement in the body due to binding to lipophilic
sites, whereas very hydrophilic molecules can be trapped in water
pockets [28]. In the specialized literature it has been noted that the
biological effect of some substances is significantly reduced with an
increase in their solubility in water. The competition of these two
processes leads to the existence of some optimal (in the region of
the maximum) hydrophobic properties of chemical compounds. In
the range of values n = 9 – 14 (additional hydrophobicity of the base
molecule is equal to Δπ ≈ 5 – 7 conventional units), the toxicity of the
drugs reaches its maximum value. Their maximum radioprotective
effect is also manifested in this same range of n index values. Thus,
although chemical compounds with index n > 14 have values of Z
< (2.706 – 2.715) CU, as well as dH < (-0.069 – 0.0922) bits and
are probably potentially active radioprotectors, nevertheless, their
radioprotective effect is limited by the hydrophobic properties of
the molecules.
Figures 2A and 2B demonstrate the change in toxicity (1/ LD50) of agents and the radioprotective effect (A, %) of chemical compounds of a series of N-alkyl-substituted Bunte salts R-NHCH2CH2SSO3H depending on their hydrophobic properties; substituent R = CH3(CH2)n, where n = 0, 1, …, 17. However, instead of the analytical approximation (22) commonly used in the literature, it is proposed here to use a more realistic approximation of the relationship between bioactivity and the hydrophobic properties of molecules [29]. The use of equation (22) can lead, for some values of the parameter Δπ, to negative values of biological activity. Therefore, it is preferable to describe the dependence of the toxicity of molecules on their hydrophobic properties using a more realistic nonlinear approximation, which does not have the indicated drawback (Figure 2A):



The approximation statistics (23) will be as follows: a1 = 0.110 ± 0.011, b1 = 5.65 ± 0.109, c1 = -1.43 ± 0.185, d1 = 0.00216 ± 0.00056, (RMSE = 0.0156), N = 18; R2 = 0.94 is the coefficient of determination. Therefore, 94% of the total scatter can be explained by regression (23) and only 6% can be explained by some unaccounted factors. Such a relationship of signs can be attributed to a causal relationship [30]. The empirical value of the F-ratio is F =12.0 > F0.05cr (f1= 1; f2 =17)=4.54 . The above statistics indicate that the null hypothesis of independence of the explanatory feature Δπ and the bio response (1/LD50) can be rejected. Equations (22) and (23) indicate that there is a range of values of the index n for which the toxicity of molecules is maximal (Figure 2A). A similar equation was obtained for the relationship between RPA and the hydrophobicity of molecules (A, %) (Figure 2B):


The toxicity of molecules has a maximum (Figure 2A) in the range of values Δπ ≈ 5 − 7 conventional units (i.e. for n ≈ 10 – 13). The maximum radioprotective activity (Figure 2B) is in the range of values Δπ =1− 2 conventional units (n = 2 − 4) . It is well known that hydrophobicity characterizes the ability of a substance to dissolve in water. At the same time, the property of lipophilicity promotes the dissolution of a chemical compound in fats, oils, lipids (penetrate through cell membranes), non-polar solvents, that is, influence the pharmacokinetic and pharmacodynamic stages of the substance’s behavior in the body. A sharp decrease in the radioprotective activity of molecules from 90% to zero when moving from n = 7 to n = 8 (Table 4) may also be associated with the instability of the linearity of the flexible hydrocarbon chain with an increase in the number of CH2 groups (links) and the folding of the molecule into a spiral or ball.
Table 4: Additional hydrophobic contribution (Δπn), as well as electronic and informational features of chemical compounds of the CH3(CH2)nNHCH 2CH2SSO3H series.

The longer the chain, the more flexibility it has. σ-Chemical bonds between the links are axisymmetric and rotation around them does not require significant energy expenditure. The flexibility of a linear molecule means that the molecule can transform into an irregular coil (a persistent or uniformly elastic model). The molecule’s memory of its linearity decreases with an increase in the number of links (very roughly estimated at a minimum value of 4 – 5). In many cases, the thermodynamic advantage of the formation of aggregates of hydrophobic groups (CH2) with a density sufficient to displace water molecules turns out to be the leading factor in stabilizing the coil [31]. It is the conformational properties of molecules that determine the biological characteristics of molecules [32]. It should also be noted that the formation of the ball depends significantly on the pH of the aqueous medium, which changes significantly after intense irradiation.
Conclusion
The performed factor, correlation and cluster analyses demonstrated the possibility of using molecular features to predict the effective radioprotective activity of sulfur-containing chemical compounds. The demonstrated ability to use molecular signatures derived from knowledge of only the molecular structure of an agent can greatly facilitate the screening of new drugs. The statistically significant relationships between the electronic and informational features for effective radioprotectors and for inactive agents can serve as an additional feature for separating chemical compounds according to their radioprotective activity. That is, there is a structural shift in the relationships of molecular features. Such differences with a structural shift in relationships also occur for other types of bioactivities [33].
Sigmoid approach on AIBL case study
None.
References
- Abarenkov IV, Brattsev VF, Tulub AI(1989) Principles of Quantum Chemistry.Moscow, Vysshaya shkola 303.
- Brandt NB, Chudinov SM(1980) Energy spectra of electrons and phonons in metals.Moscow State University,Moscow PP. 344.
- Veljkovič V, Lalovič D (1975) Superconductivity and the Periodic System. Phys Rev B 11(11): 4242-4244.
- Quastler H (1964) The Emergence of Biological Organization. New Haven and London, Uhle University Press.
- Kolmogorov AN (1987) Information Theory and Theory of Algorithms. Moscow, Nauka PP. 303.
- Alexander P, Bacq ZM, Causens SF, Causins SF, Fox M, et al. (1955) Mode of Action of Some Substances Which Protect against the Lethal Effects of X-Rays. Radiat Res 2(4): 392-415.
- Sweeney TR (1979) A Survey of Compounds from the Antiradiation Drug Development Program of the US Army Medical Research and Development Command. Walter Reed Army Institute of Research. Washington, D.C.
- Romantsev EF (1968) Radiation and chemical protection. Moscow PP. 248.
- Yashunsky VG, Kovtun VYu (1985) New chemical means of protection against ionizing radiation. Advances in Chemistry 54(1): 126-161.
- Yashunsky VG (1975) Advances in the search for chemical radioprotectors. Advances in Chemistry 44(3): 531-574.
- Carroll FI, Wall ME (1970) N-Substituted Aminoeyhanethiols and N-Substituted Aminoethanethiols-Sulforic Acids as Radioprotective Agents. Journal of Pharmaceutical Sciences 59(9): 1350-1352.
- Tank LI (1959) Collection of Proceedings of the Scientific Conference (December 1957). State Medical Publishing House, Kyiv PP. 260-269.
- Kobzar AI (2016) Applied Mathematical Statistics. For Engineers and Scientists. Moscow, Fizmatlit. PP. 816.
- Sachs L (1972) Statistishe Ausweryungsmethoden. Springer-Werlag. Berlin, Heidelberg, New York, USA.
- Förster E, Rönz B (1979) Metohden der Korrelations- und Regressionsanalyse. Verlag Die Wirtschaft, Berlin.
- Radiation Research (1967) Proceedings of the Third International Congress of Radiation Research held at Cortina d’Ampezzo, Italy. North Holland Publishing Company, Amsterdam.
- Mukhomorov VK (2011) Bipolarons Structure Properties. LAP Lambert Academic Publishing. Saarbrü Deutchland. PP. 278.
- Aldenderfer MS, Blashfield RK (1984) Cluster Analysis. Series: Quantative Applications in the Social Sciences. 44. San Francisco. W.H. Freeman. Sage University Paper. Bewerly Hills. London, NewDelhi PP.88.
- Gitis L Kh (2003) Statistical classification and cluster analysis. Moscow State Mining University, Moscow PP. 157.
- Steinhaus H (1956) Sur la division des corps materiels en parties. Bull Acad Polon Sci PP. 801-804.
- Vuchkov I, Bayadzhieva L, Solakov E (1987) Applied linear regression analysis. PP. 239.
- (1989) Handbook of Applicable Mathematics. University of Lancaster. John Wiley & Sons. Chichester-New York PP. 526.
- Romanovsky VI (1947) Application of statistics in experimental work. OGIZ, Moscow-Leningrad PP. 247.
- D'Agostino RB (1970) Linear estimation of the normal distribution standard deviation. The Amer Statist 24(6): 14-15.
- Zaitsev GN (1990) Mathematics in experimental botany. Moscow Science PP. 295.
- Mukhomorov VK (2014) Biological Activity of Chemical Compounds and Their Molecular Structure - Information Approach. Journal Chem Eng and Chem Res 1(1): 54-65.
- Suvorov NN, Shashkov VS (1975) Chemistry and pharmacology of means of preventing radiation injuries. Moscow. Atomizdat PP. 224.
- Leo A, Hansch C, Elkins D (1971) Partition coefficients and their uses. Chem Rev 71(6): 525-616.
- Hansch C (1980) On the use of quantitative structure-activity relationships in drug design. Chemical and Pharmaceutical Journal 14(10): 15-30.
- Mukhomorov VK (2011) Entropy Approach to the Study of Biological Activity of Chemical Compounds. Advances in Biological Chemistry 1(1): 1-5.
- Draper NR, Smith H (1986) Applied Regression Analysis. Part 2. John Willey & Sons. New York – Chichester. 366.
- Grosberg AYu, Khokhlov AR (1989) Statistical physics of macromolecules. Moscow. Main editorial office of physical and mathematical literature PP. 237.
- Mukhomorov VK (2021) Statistical modeling of bioactivity of chemical compounds. Structure and properties. LAP Lambert Academic Publishing RU Part 1 PP. 308. Part 2 PP. 380.
-
Mukhomorov VK*. Multidimensional Classification of Radioprotective Activity of Sulfur-Containing Chemical Agents. Annal Biostat & Biomed Appli. 6(2): 2024. ABBA.MS.ID.000635.
Modeling; cluster analysis; molecular features; pseudopotential; electronic feature; information function; structural shift; dose; classification rule
-

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.






